[English] 日本語
 Yorodumi
Yorodumi- EMDB-19962: Archaellum filament from the Halobacterium salinarum deltaAgl27 strain -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Archaellum filament from the Halobacterium salinarum deltaAgl27 strain | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | archaellum / haloarcheon / archaellin / STRUCTURAL PROTEIN | |||||||||
| Biological species |  Halobacterium salinarum (Halophile) Halobacterium salinarum (Halophile) | |||||||||
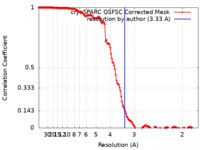
| Method | helical reconstruction / cryo EM / Resolution: 3.33 Å | |||||||||
 Authors Authors | Grossman-Haham I / Shahar A | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Perturbed N-glycosylation of Halobacterium salinarum archaellum filaments leads to filament bundling and compromised cell motility Authors: Mashni L / Vershinin Z / Zalk R / Shahar A / Eichler J / Grossman-Haham I | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19962.map.gz emd_19962.map.gz | 141.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19962-v30.xml emd-19962-v30.xml emd-19962.xml emd-19962.xml | 15.8 KB 15.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19962_fsc.xml emd_19962_fsc.xml | 13.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_19962.png emd_19962.png | 162.3 KB | ||
| Filedesc metadata |  emd-19962.cif.gz emd-19962.cif.gz | 5.7 KB | ||
| Others |  emd_19962_additional_1.map.gz emd_19962_additional_1.map.gz emd_19962_half_map_1.map.gz emd_19962_half_map_1.map.gz emd_19962_half_map_2.map.gz emd_19962_half_map_2.map.gz | 233.9 MB 255.2 MB 255.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19962 http://ftp.pdbj.org/pub/emdb/structures/EMD-19962 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19962 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19962 | HTTPS FTP |
-Validation report
| Summary document |  emd_19962_validation.pdf.gz emd_19962_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_19962_full_validation.pdf.gz emd_19962_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_19962_validation.xml.gz emd_19962_validation.xml.gz | 22.9 KB | Display | |
| Data in CIF |  emd_19962_validation.cif.gz emd_19962_validation.cif.gz | 29.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19962 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19962 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19962 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19962 | HTTPS FTP |
-Related structure data
| Related structure data |  9etuMC  9eq7C  9esmC M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_19962.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19962.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.89 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_19962_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_19962_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_19962_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Archaellum filament
| Entire | Name: Archaellum filament |
|---|---|
| Components |
|
-Supramolecule #1: Archaellum filament
| Supramolecule | Name: Archaellum filament / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Halobacterium salinarum (Halophile) Halobacterium salinarum (Halophile) |
| Molecular weight | Theoretical: 20.466 KDa |
-Macromolecule #1: Archaellin
| Macromolecule | Name: Archaellin / type: protein_or_peptide / ID: 1 Details: Since Hbt. salinarum encodes five archaellins (i.e., FlaA1, FlaA2, FlaB1, FlaB2, and FlaB3) and their arrangement within the archaellum filaments is unknown, we further refined the cryo-EM ...Details: Since Hbt. salinarum encodes five archaellins (i.e., FlaA1, FlaA2, FlaB1, FlaB2, and FlaB3) and their arrangement within the archaellum filaments is unknown, we further refined the cryo-EM map without applying symmetry, in an attempt to resolve the positions of these archaellins within the filament, as done previously with reconstruction of the Methanocaldococcus villosus archaellum. Symmetry-free refinement improved the overall resolution map to 3.1 A and revealed differences in density among archaellin subunits. Nonetheless, we were unable to identify features that would allow us to unambiguously assign specific archaellins into the density, perhaps because the regions that distinguish each archaellin are few, short, and mostly predicted to lack defined secondary structure, or because the five archaellins are not organized in a repeating pattern. Consequently, we built a model into the central region of the cryo-EM map comprising 26 archaellin subunits that share a consensus sequence, in which identical residues among the five archaellins are explicitly modelled, with those variable residues usually being modelled as alanine residues (UNK). Number of copies: 25 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Halobacterium salinarum (Halophile) Halobacterium salinarum (Halophile) |
| Molecular weight | Theoretical: 20.107971 KDa |
| Sequence | String: MFEFITDEDE RGQVGIGTLI VFIAMVLVAA IAAGVLINTA GYLQSKGSAT GEEASAQVSN RINIVSAYGN V(UNK) (UNK)(UNK)(UNK)VDYV NLTVRQAAGA DNINL(UNK)KSTI QWIGPD(UNK)ATT LTY(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK) ...String: MFEFITDEDE RGQVGIGTLI VFIAMVLVAA IAAGVLINTA GYLQSKGSAT GEEASAQVSN RINIVSAYGN V(UNK) (UNK)(UNK)(UNK)VDYV NLTVRQAAGA DNINL(UNK)KSTI QWIGPD(UNK)ATT LTY(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)ENFTT(UNK)S IKG(UNK)(UNK)(UNK)(UNK)VLV DQSDRIKVIM YA(UNK) (UNK)V(UNK)(UNK)(UNK)L (UNK)(UNK)G(UNK)EVQLTV TTQYGSKTTY WAQVPESLKD KNAV(UNK)L |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: OTHER / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)