[English] 日本語
 Yorodumi
Yorodumi- EMDB-19910: Cyanide dihydratase from Bacillus pumilus C1 E155R variant with a... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cyanide dihydratase from Bacillus pumilus C1 E155R variant with altered helical twist. | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Left-handed helix. / HYDROLASE | |||||||||
| Function / homology | Nitrilase/Cyanide hydratase / Nitrilases / cyanide hydratase signature 1. / Nitrilase/cyanide hydratase, conserved site / nitrilase activity / Carbon-nitrogen hydrolase superfamily / Carbon-nitrogen hydrolase / Carbon-nitrogen hydrolase domain profile. / Carbon-nitrogen hydrolase / Cyanide dihydratase Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Dlamini LS / Woodward JD / Sewell BT | |||||||||
| Funding support |  South Africa, 1 items South Africa, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cyanide dihydratase from Bacillus pumilus C1 E155R variant with altered helical twist Authors: Dlamini LS / Woodward JD / Sewell BT | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19910.map.gz emd_19910.map.gz | 166.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19910-v30.xml emd-19910-v30.xml emd-19910.xml emd-19910.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19910_fsc.xml emd_19910_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_19910.png emd_19910.png | 48.4 KB | ||
| Filedesc metadata |  emd-19910.cif.gz emd-19910.cif.gz | 6 KB | ||
| Others |  emd_19910_half_map_1.map.gz emd_19910_half_map_1.map.gz emd_19910_half_map_2.map.gz emd_19910_half_map_2.map.gz | 139.6 MB 139.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19910 http://ftp.pdbj.org/pub/emdb/structures/EMD-19910 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19910 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19910 | HTTPS FTP |
-Validation report
| Summary document |  emd_19910_validation.pdf.gz emd_19910_validation.pdf.gz | 1.2 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_19910_full_validation.pdf.gz emd_19910_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  emd_19910_validation.xml.gz emd_19910_validation.xml.gz | 20.1 KB | Display | |
| Data in CIF |  emd_19910_validation.cif.gz emd_19910_validation.cif.gz | 26.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19910 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19910 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19910 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19910 | HTTPS FTP |
-Related structure data
| Related structure data |  9er3MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_19910.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19910.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.825 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_19910_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
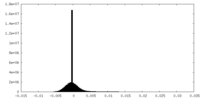
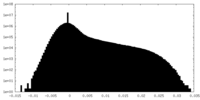
| Density Histograms |
-Half map: #1
| File | emd_19910_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
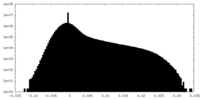
| Density Histograms |
- Sample components
Sample components
-Entire : Cyanide dihydratase from Bacillus pumilus C1 E155R variant with a...
| Entire | Name: Cyanide dihydratase from Bacillus pumilus C1 E155R variant with altered helical twist. |
|---|---|
| Components |
|
-Supramolecule #1: Cyanide dihydratase from Bacillus pumilus C1 E155R variant with a...
| Supramolecule | Name: Cyanide dihydratase from Bacillus pumilus C1 E155R variant with altered helical twist. type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 703 KDa |
-Macromolecule #1: Cyanide dihydratase
| Macromolecule | Name: Cyanide dihydratase / type: protein_or_peptide / ID: 1 / Number of copies: 19 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 36.19873 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: TSIYPKFRAA AVQAAPIYLN LEASVEKSCE LIDEAASNGA KLVAFPEAFL PGYPWFAFIG HPEYTRKFYH ELYKNAVEIP SLAIQKISE AAKRNETYVC ISCSEKDGGS LYLAQLWFNP NGDLIGKHRK MRASVAERLI WGDGSGSMMP VFQTRIGNLG G LMCWEHQV ...String: TSIYPKFRAA AVQAAPIYLN LEASVEKSCE LIDEAASNGA KLVAFPEAFL PGYPWFAFIG HPEYTRKFYH ELYKNAVEIP SLAIQKISE AAKRNETYVC ISCSEKDGGS LYLAQLWFNP NGDLIGKHRK MRASVAERLI WGDGSGSMMP VFQTRIGNLG G LMCWEHQV PLDLMAMNAQ NEQVHVASWP GYFDDEISSR YYAIATQTFV LMTSSIYTEE MKEMICLTQE QRDYFETFKS GH TCIYGPD GEPISDMVPA ETEGIAYAEI DVERVIDYKY YIDPAGHYSN QSLSMNFNQQ PTPVVKHLNH QKNEVFTYED IQ UniProtKB: Cyanide dihydratase |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 1.8 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Component - Concentration: 50.0 mM / Component - Formula: Tris-HCl / Component - Name: Tris-(hydroxymethyl) aminomethane / Details: 50 mM NaCL, 50 mM Tris-HCl |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: OTHER / Pretreatment - Pressure: 2.0 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK I |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 31418 / Average exposure time: 2.04 sec. / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: Other / Chain - Initial model type: other Details: The initial model was built ab initio using the map and amino acid sequence. |
|---|---|
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 101.248 / Target criteria: Cross-correlation coefficient |
| Output model |  PDB-9er3: |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)