+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of UVSSA(VHS)-CSA-DDB1-DDA1 | |||||||||
 Map data Map data | Composite map of UVSSA(VHS)-CSA-DDB1-DDA1 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ubiquitin ligase / DNA repair / LIGASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationRNA polymerase inhibitor activity / nucleotide-excision repair complex / regulation of transcription-coupled nucleotide-excision repair / response to auditory stimulus / single strand break repair / double-strand break repair via classical nonhomologous end joining / positive regulation by virus of viral protein levels in host cell / chromatin-protein adaptor activity / spindle assembly involved in female meiosis / epigenetic programming in the zygotic pronuclei ...RNA polymerase inhibitor activity / nucleotide-excision repair complex / regulation of transcription-coupled nucleotide-excision repair / response to auditory stimulus / single strand break repair / double-strand break repair via classical nonhomologous end joining / positive regulation by virus of viral protein levels in host cell / chromatin-protein adaptor activity / spindle assembly involved in female meiosis / epigenetic programming in the zygotic pronuclei / UV-damage excision repair / biological process involved in interaction with symbiont / regulation of mitotic cell cycle phase transition / WD40-repeat domain binding / Cul4A-RING E3 ubiquitin ligase complex / Cul4-RING E3 ubiquitin ligase complex / Cul4B-RING E3 ubiquitin ligase complex / ubiquitin ligase complex scaffold activity / negative regulation of reproductive process / negative regulation of developmental process / RNA polymerase II complex binding / viral release from host cell / cullin family protein binding / response to X-ray / ectopic germ cell programmed cell death / positive regulation of viral genome replication / response to UV / protein autoubiquitination / ubiquitin-like ligase-substrate adaptor activity / site of DNA damage / transcription-coupled nucleotide-excision repair / proteasomal protein catabolic process / positive regulation of gluconeogenesis / positive regulation of DNA repair / nucleotide-excision repair / sperm end piece / Recognition of DNA damage by PCNA-containing replication complex / regulation of circadian rhythm / DNA Damage Recognition in GG-NER / Dual Incision in GG-NER / Transcription-Coupled Nucleotide Excision Repair (TC-NER) / Formation of TC-NER Pre-Incision Complex / Wnt signaling pathway / nuclear matrix / Formation of Incision Complex in GG-NER / protein polyubiquitination / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / positive regulation of protein catabolic process / cellular response to UV / rhythmic process / positive regulation of proteasomal ubiquitin-dependent protein catabolic process / chromosome / site of double-strand break / sperm principal piece / Neddylation / response to oxidative stress / sperm midpiece / ubiquitin-dependent protein catabolic process / damaged DNA binding / perikaryon / proteasome-mediated ubiquitin-dependent protein catabolic process / protein-macromolecule adaptor activity / chromosome, telomeric region / protein ubiquitination / DNA repair / apoptotic process / DNA damage response / negative regulation of apoptotic process / protein-containing complex binding / nucleolus / protein-containing complex / : / DNA binding / extracellular exosome / zinc ion binding / nucleoplasm / nucleus / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Lee S-H / Sixma TK | |||||||||
| Funding support |  Netherlands, European Union, 2 items Netherlands, European Union, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: The small CRL4 ubiquitin ligase component DDA1 regulates transcription-coupled repair dynamics. Authors: Diana A Llerena Schiffmacher / Shun-Hsiao Lee / Katarzyna W Kliza / Arjan F Theil / Masaki Akita / Angela Helfricht / Karel Bezstarosti / Camila Gonzalo-Hansen / Haico van Attikum / Matty ...Authors: Diana A Llerena Schiffmacher / Shun-Hsiao Lee / Katarzyna W Kliza / Arjan F Theil / Masaki Akita / Angela Helfricht / Karel Bezstarosti / Camila Gonzalo-Hansen / Haico van Attikum / Matty Verlaan-de Vries / Alfred C O Vertegaal / Jan H J Hoeijmakers / Jurgen A Marteijn / Hannes Lans / Jeroen A A Demmers / Michiel Vermeulen / Titia K Sixma / Tomoo Ogi / Wim Vermeulen / Alex Pines /     Abstract: Transcription-blocking DNA lesions are specifically targeted by transcription-coupled nucleotide excision repair (TC-NER), which removes a broad spectrum of DNA lesions to preserve transcriptional ...Transcription-blocking DNA lesions are specifically targeted by transcription-coupled nucleotide excision repair (TC-NER), which removes a broad spectrum of DNA lesions to preserve transcriptional output and thereby cellular homeostasis to counteract aging. TC-NER is initiated by the stalling of RNA polymerase II at DNA lesions, which triggers the assembly of the TC-NER-specific proteins CSA, CSB and UVSSA. CSA, a WD40-repeat containing protein, is the substrate receptor subunit of a cullin-RING ubiquitin ligase complex composed of DDB1, CUL4A/B and RBX1 (CRL4). Although ubiquitination of several TC-NER proteins by CRL4 has been reported, it is still unknown how this complex is regulated. To unravel the dynamic molecular interactions and the regulation of this complex, we apply a single-step protein-complex isolation coupled to mass spectrometry analysis and identified DDA1 as a CSA interacting protein. Cryo-EM analysis shows that DDA1 is an integral component of the CRL4 complex. Functional analysis reveals that DDA1 coordinates ubiquitination dynamics during TC-NER and is required for efficient turnover and progression of this process. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18398.map.gz emd_18398.map.gz | 92.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18398-v30.xml emd-18398-v30.xml emd-18398.xml emd-18398.xml | 27.9 KB 27.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_18398.png emd_18398.png | 50.3 KB | ||
| Filedesc metadata |  emd-18398.cif.gz emd-18398.cif.gz | 9.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18398 http://ftp.pdbj.org/pub/emdb/structures/EMD-18398 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18398 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18398 | HTTPS FTP |
-Related structure data
| Related structure data |  8qh5MC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18398.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18398.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Composite map of UVSSA(VHS)-CSA-DDB1-DDA1 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.057 Å | ||||||||||||||||||||||||||||||||||||
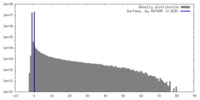
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Ternary complex of UVSSA(VHS)-CSA-DDB1-DDA1
| Entire | Name: Ternary complex of UVSSA(VHS)-CSA-DDB1-DDA1 |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of UVSSA(VHS)-CSA-DDB1-DDA1
| Supramolecule | Name: Ternary complex of UVSSA(VHS)-CSA-DDB1-DDA1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #2-#4, #1 Details: Ternary complex of ubiquitinated UVSSA-USP7-CSA-DDB1-DDA1. USP7 is invisible in the cryoEM map. |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 210 KDa |
-Macromolecule #1: UV-stimulated scaffold protein A
| Macromolecule | Name: UV-stimulated scaffold protein A / type: protein_or_peptide / ID: 1 / Details: The construct contains an N-terminal His tag. / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 82.923164 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAHHHHHHSA ALEVLFQGPG MDQKLSKLVE ELTTSGEPRL NPEKMKELKK ICKSSEEQLS RAYRLLIAQL TQEHAEIRLS AFQIVEELF VRSHQFRMLV VSNFQEFLEL TLGTDPAQPL PPPREAAQRL RQATTRAVEG WNEKFGEAYK KLALGYHFLR H NKKVDFQD ...String: MAHHHHHHSA ALEVLFQGPG MDQKLSKLVE ELTTSGEPRL NPEKMKELKK ICKSSEEQLS RAYRLLIAQL TQEHAEIRLS AFQIVEELF VRSHQFRMLV VSNFQEFLEL TLGTDPAQPL PPPREAAQRL RQATTRAVEG WNEKFGEAYK KLALGYHFLR H NKKVDFQD TNARSLAERK REEEKQKHLD KIYQERASQA EREMQEMSGE IESCLTEVES CFRLLVPFDF DPNPETESLG MA SGMSDAL RSSCAGQVGP CRSGTPDPRD GEQPCCSRDL PASAGHPRAG GGAQPSQTAT GDPSDEDEDS DLEEFVRSHG LGS HKYTLD VELCSEGLKV QENEDNLALI HAARDTLKLI RNKFLPAVCS WIQRFTRVGT HGGCLKRAID LKAELELVLR KYKE LDIEP EGGERRRTEA LGDAEEDEDD EDFVEVPEKE GYEPHIPDHL RPEYGLEAAP EKDTVVRCLR TRTRMDEEVS DPTSA AAQL RQLRDHLPPP SSASPSRALP EPQEAQKLAA ERARAPVVPY GVDLHYWGQE LPTAGKIVKS DSQHRFWKPS EVEEEV VNA DISEMLRSRH ITFAGKFEPV QHWCRAPRPD GRLCERQDRL KCPFHGKIVP RDDEGRPLDP EDRAREQRRQ LQKQERP EW QDPELMRDVE AATGQDLGSS RYSGKGRGKK RRYPSLTNLK AQADTARARI GRKVFAKAAV RRVVAAMNRM DQKKHEKF S NQFNYALN UniProtKB: UV-stimulated scaffold protein A |
-Macromolecule #2: DNA excision repair protein ERCC-8
| Macromolecule | Name: DNA excision repair protein ERCC-8 / type: protein_or_peptide / ID: 2 Details: The construct contains a Strep tag II at the C-terminus Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 45.465613 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLGFLSARQT GLEDPLRLRR AESTRRVLGL ELNKDRDVER IHGGGINTLD IEPVEGRYML SGGSDGVIVL YDLENSSRQS YYTCKAVCS IGRDHPDVHR YSVETVQWYP HDTGMFTSSS FDKTLKVWDT NTLQTADVFN FEETVYSHHM SPVSTKHCLV A VGTRGPKV ...String: MLGFLSARQT GLEDPLRLRR AESTRRVLGL ELNKDRDVER IHGGGINTLD IEPVEGRYML SGGSDGVIVL YDLENSSRQS YYTCKAVCS IGRDHPDVHR YSVETVQWYP HDTGMFTSSS FDKTLKVWDT NTLQTADVFN FEETVYSHHM SPVSTKHCLV A VGTRGPKV QLCDLKSGSC SHILQGHRQE ILAVSWSPRY DYILATASAD SRVKLWDVRR ASGCLITLDQ HNGKKSQAVE SA NTAHNGK VNGLCFTSDG LHLLTVGTDN RMRLWNSSNG ENTLVNYGKV CNNSKKGLKF TVSCGCSSEF VFVPYGSTIA VYT VYSGEQ ITMLKGHYKT VDCCVFQSNF QELYSGSRDC NILAWVPSLY EPVPDDDETT TKSQLNPAFE DAWSSSDEEG GTSA WSHPQ FEK UniProtKB: DNA excision repair protein ERCC-8 |
-Macromolecule #3: DNA damage-binding protein 1
| Macromolecule | Name: DNA damage-binding protein 1 / type: protein_or_peptide / ID: 3 / Details: The construct contains an N-terminal His tag. / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 129.298867 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAHHHHHHSA ALEVLFQGPG MSYNYVVTAQ KPTAVNGCVT GHFTSAEDLN LLIAKNTRLE IYVVTAEGLR PVKEVGMYGK IAVMELFRP KGESKDLLFI LTAKYNACIL EYKQSGESID IITRAHGNVQ DRIGRPSETG IIGIIDPECR MIGLRLYDGL F KVIPLDRD ...String: MAHHHHHHSA ALEVLFQGPG MSYNYVVTAQ KPTAVNGCVT GHFTSAEDLN LLIAKNTRLE IYVVTAEGLR PVKEVGMYGK IAVMELFRP KGESKDLLFI LTAKYNACIL EYKQSGESID IITRAHGNVQ DRIGRPSETG IIGIIDPECR MIGLRLYDGL F KVIPLDRD NKELKAFNIR LEELHVIDVK FLYGCQAPTI CFVYQDPQGR HVKTYEVSLR EKEFNKGPWK QENVEAEASM VI AVPEPFG GAIIIGQESI TYHNGDKYLA IAPPIIKQST IVCHNRVDPN GSRYLLGDME GRLFMLLLEK EEQMDGTVTL KDL RVELLG ETSIAECLTY LDNGVVFVGS RLGDSQLVKL NVDSNEQGSY VVAMETFTNL GPIVDMCVVD LERQGQGQLV TCSG AFKEG SLRIIRNGIG IHEHASIDLP GIKGLWPLRS DPNRETDDTL VLSFVGQTRV LMLNGEEVEE TELMGFVDDQ QTFFC GNVA HQQLIQITSA SVRLVSQEPK ALVSEWKEPQ AKNISVASCN SSQVVVAVGR ALYYLQIHPQ ELRQISHTEM EHEVAC LDI TPLGDSNGLS PLCAIGLWTD ISARILKLPS FELLHKEMLG GEIIPRSILM TTFESSHYLL CALGDGALFY FGLNIET GL LSDRKKVTLG TQPTVLRTFR SLSTTNVFAC SDRPTVIYSS NHKLVFSNVN LKEVNYMCPL NSDGYPDSLA LANNSTLT I GTIDEIQKLH IRTVPLYESP RKICYQEVSQ CFGVLSSRIE VQDTSGGTTA LRPSASTQAL SSSVSSSKLF SSSTAPHET SFGEEVEVHN LLIIDQHTFE VLHAHQFLQN EYALSLVSCK LGKDPNTYFI VGTAMVYPEE AEPKQGRIVV FQYSDGKLQT VAEKEVKGA VYSMVEFNGK LLASINSTVR LYEWTTEKEL RTECNHYNNI MALYLKTKGD FILVGDLMRS VLLLAYKPME G NFEEIARD FNPNWMSAVE ILDDDNFLGA ENAFNLFVCQ KDSAATTDEE RQHLQEVGLF HLGEFVNVFC HGSLVMQNLG ET STPTQGS VLFGTVNGMI GLVTSLSESW YNLLLDMQNR LNKVIKSVGK IEHSFWRSFH TERKTEPATG FIDGDLIESF LDI SRPKMQ EVVANLQYDD GSGMKREATA DDLIKVVEEL TRIH UniProtKB: DNA damage-binding protein 1 |
-Macromolecule #4: DET1- and DDB1-associated protein 1
| Macromolecule | Name: DET1- and DDB1-associated protein 1 / type: protein_or_peptide / ID: 4 Details: The construct contains a TwinStrep tag and a Flag tag at the C-terminus. Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 16.997615 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MADFLKGLPV YNKSNFSRFH ADSVCKASNR RPSVYLPTRE YPSEQIIVTE KTNILLRYLH QQWDKKNAAK KRDQEQVELE GESSAPPRK VARTDSPDMH EDTDVLFQGP GAWSHPQFEK GGGSGGGSGG GSWSHPQFEK GASGEDYKDD DDK UniProtKB: DET1- and DDB1-associated protein 1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.13 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Details: The grid was coated with graphene oxide. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV | ||||||||||||
| Details | This sample was glutaraldehyde crosslinked |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Details | Collected on Krios 1 at Netherlands Center for Electron Nanoscopy (NeCEN) |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 1382 / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Details | Additional densities are observed near several cysteine residues (A/Cys222, A/Cys260, A/Cys288, B/Cys363, B/Cys725, B/Cys1008). We expect these are oxidized products or crosslinking side products. | ||||||||||||||||||
| Refinement | Space: REAL / Protocol: RIGID BODY FIT | ||||||||||||||||||
| Output model |  PDB-8qh5: |
 Movie
Movie Controller
Controller












 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)