+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Composite map of the Gallus gallus 80S non-rotated ribosome | |||||||||
 Map data Map data | Composite map of the Gallus gallus 80S non-rotated ribosome | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | translation / ribosome | |||||||||
| Function / homology |  Function and homology information Function and homology informationTNFR1-mediated ceramide production / Major pathway of rRNA processing in the nucleolus and cytosol / Protein hydroxylation / Translesion synthesis by REV1 / Recognition of DNA damage by PCNA-containing replication complex / Translesion Synthesis by POLH / Activation of NF-kappaB in B cells / Oxygen-dependent proline hydroxylation of Hypoxia-inducible Factor Alpha / Spry regulation of FGF signaling / Downregulation of ERBB2:ERBB3 signaling ...TNFR1-mediated ceramide production / Major pathway of rRNA processing in the nucleolus and cytosol / Protein hydroxylation / Translesion synthesis by REV1 / Recognition of DNA damage by PCNA-containing replication complex / Translesion Synthesis by POLH / Activation of NF-kappaB in B cells / Oxygen-dependent proline hydroxylation of Hypoxia-inducible Factor Alpha / Spry regulation of FGF signaling / Downregulation of ERBB2:ERBB3 signaling / APC/C:Cdc20 mediated degradation of Cyclin B / Autodegradation of Cdh1 by Cdh1:APC/C / SCF-beta-TrCP mediated degradation of Emi1 / APC/C:Cdc20 mediated degradation of Securin / APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 / Cdc20:Phospho-APC/C mediated degradation of Cyclin A / APC-Cdc20 mediated degradation of Nek2A / EGFR downregulation / SCF(Skp2)-mediated degradation of p27/p21 / Degradation of beta-catenin by the destruction complex / TCF dependent signaling in response to WNT / Downstream TCR signaling / NRIF signals cell death from the nucleus / p75NTR recruits signalling complexes / NF-kB is activated and signals survival / Activated NOTCH1 Transmits Signal to the Nucleus / Downregulation of SMAD2/3:SMAD4 transcriptional activity / SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription / Senescence-Associated Secretory Phenotype (SASP) / FCERI mediated NF-kB activation / Regulation of innate immune responses to cytosolic DNA / Autodegradation of the E3 ubiquitin ligase COP1 / RAD18 and ubiquitinated PCNA-mediated recruitment of translesion polymerases / Nucleotide Excision Repair / Deactivation of the beta-catenin transactivating complex / TRAF6 mediated induction of proinflammatory cytokines / TAK1 activates NFkB by phosphorylation and activation of IKKs complex / NFkB activation mediated by RIP1 complexed with activated TLR3 / Activated TAK1 mediates p38 MAP kinase phosphorylation / Activated TAK1 mediates Jun kinases (JNK) phosphorylation and activation / activated TAK1 mediates p38 MAPK activation / JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 / AUF1 (hnRNP D0) binds and destabilizes mRNA / Degradation of AXIN / Degradation of DVL / Regulation of FZD by ubiquitination / N-glycan trimming in the ER and Calnexin/Calreticulin cycle / Regulation of TNFR1 signaling / TNFR1-induced NF-kappa-B signaling pathway / Hedgehog ligand biogenesis / CLEC7A (Dectin-1) signaling / Degradation of GLI1 by the proteasome / GLI3 is processed to GLI3R by the proteasome / Hedgehog 'on' state / Negative regulation of FGFR1 signaling / Negative regulation of FGFR2 signaling / Negative regulation of FGFR3 signaling / Negative regulation of FGFR4 signaling / Translesion synthesis by POLK / Translesion synthesis by POLI / Termination of translesion DNA synthesis / TNFR2 non-canonical NF-kB pathway / Negative regulation of MAPK pathway / Regulation of necroptotic cell death / MAP3K8 (TPL2)-dependent MAPK1/3 activation / HDR through Homologous Recombination (HRR) / MAPK6/MAPK4 signaling / Josephin domain DUBs / Ovarian tumor domain proteases / Formation of Incision Complex in GG-NER / Gap-filling DNA repair synthesis and ligation in GG-NER / Dual Incision in GG-NER / NFkB and MAPK activation mediated by TRAF6 upon TLR7 or TLR21 stimulation / Formation of TC-NER Pre-Incision Complex / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / Regulation of TP53 Activity through Phosphorylation / Regulation of TP53 Degradation / Regulation of TP53 Activity through Methylation / Negative regulation of MET activity / Assembly of the pre-replicative complex / CDK-mediated phosphorylation and removal of Cdc6 / Ubiquitin-Mediated Degradation of Phosphorylated Cdc25A / Translation initiation complex formation / Formation of the ternary complex, and subsequently, the 43S complex / Ribosomal scanning and start codon recognition / Ubiquitin-dependent degradation of Cyclin D / PTK6 Regulates RTKs and Their Effectors AKT1 and DOK1 / FBXL7 down-regulates AURKA during mitotic entry and in early mitosis / Downregulation of ERBB2 signaling / VLDLR internalisation and degradation / Synthesis of active ubiquitin: roles of E1 and E2 enzymes / E3 ubiquitin ligases ubiquitinate target proteins / RUNX1 regulates transcription of genes involved in differentiation of HSCs / Regulation of RUNX2 expression and activity / Regulation of PTEN localization / Regulation of PTEN stability and activity / Neddylation / ER Quality Control Compartment (ERQC) / NOTCH3 Activation and Transmission of Signal to the Nucleus Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.5 Å | |||||||||
 Authors Authors | Nurullina L / Jenner L / Myasnikov A / Yusupov M | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: FEBS Lett / Year: 2024 Journal: FEBS Lett / Year: 2024Title: Cryo-EM structure of the inactive ribosome complex accumulated in chick embryo cells in cold-stress conditions. Authors: Liliia Nurullina / Salvatore Terrosu / Alexander G Myasnikov / Lasse Bohl Jenner / Marat Yusupov /    Abstract: Here, we present the high-resolution structure of the Gallus gallus 80S ribosome obtained from cold-treated chicken embryos. The translationally inactive ribosome complex contains elongation factor ...Here, we present the high-resolution structure of the Gallus gallus 80S ribosome obtained from cold-treated chicken embryos. The translationally inactive ribosome complex contains elongation factor eEF2 with GDP, SERPINE1 mRNA binding protein 1 (SERBP1) and deacylated tRNA in the P/E position, showing common features with complexes already described in mammals. Modeling of most expansion segments of G. gallus 28S ribosomal RNA allowed us to identify specific features in their structural organization and to describe areas where a marked difference between mammalian and avian ribosomes could shed light on the evolution of the expansion segments. This study provides the first structure of an avian ribosome, establishing a model for future structural and functional studies on the translational machinery in Aves. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18168.map.gz emd_18168.map.gz | 695.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18168-v30.xml emd-18168-v30.xml emd-18168.xml emd-18168.xml | 13.4 KB 13.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_18168.png emd_18168.png | 54.1 KB | ||
| Filedesc metadata |  emd-18168.cif.gz emd-18168.cif.gz | 4.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18168 http://ftp.pdbj.org/pub/emdb/structures/EMD-18168 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18168 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18168 | HTTPS FTP |
-Related structure data
| Related structure data |  8q7zMC  8q87C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18168.map.gz / Format: CCP4 / Size: 744.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18168.map.gz / Format: CCP4 / Size: 744.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Composite map of the Gallus gallus 80S non-rotated ribosome | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.803 Å | ||||||||||||||||||||||||||||||||||||
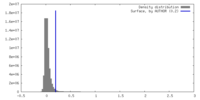
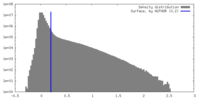
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : 80S Gallus gallus ribosome from chicken embryo
| Entire | Name: 80S Gallus gallus ribosome from chicken embryo |
|---|---|
| Components |
|
-Supramolecule #1: 80S Gallus gallus ribosome from chicken embryo
| Supramolecule | Name: 80S Gallus gallus ribosome from chicken embryo / type: complex / ID: 1 / Parent: 0 Details: 28S rRNA, 5.8S rRNA, 5S rRNA, 18S rRNA,ribosomal proteins, tRNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 4.3 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 278 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris X |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Number grids imaged: 1 / Number real images: 32439 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.3 µm / Nominal magnification: 96000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)