+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Cryo-EM structure of the DyP peroxidase-loaded encapsulin nanocompartment from Mycobacterium tuberculosis with icosahedral symmetry imposed. | |||||||||
 マップデータ マップデータ | The 2.7 A resolution cryo-EM reconstruction of Mycobacterium tuberculosis encapsulin containing DyP peroxidase | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | Encapsulin / cfp29 / nanocompartment / VIRUS LIKE PARTICLE | |||||||||
| 機能・相同性 | Type 1 encapsulin shell protein / Encapsulating protein for peroxidase / : / encapsulin nanocompartment / extracellular region / plasma membrane / Type 1 encapsulin shell protein 機能・相同性情報 機能・相同性情報 | |||||||||
| 生物種 |  Mycobacterium tuberculosis H37Rv (結核菌) Mycobacterium tuberculosis H37Rv (結核菌) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.7 Å | |||||||||
 データ登録者 データ登録者 | Willmore R / Woodward JD | |||||||||
| 資金援助 |  英国, 1件 英国, 1件
| |||||||||
 引用 引用 |  ジャーナル: To Be Published ジャーナル: To Be Publishedタイトル: The 2.7 A resolution cryo-EM reconstruction of Mycobacterium tuberculosis encapsulin nanocompartment containing DyP peroxidase. 著者: Willmore R / Woodward JD | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_18031.map.gz emd_18031.map.gz | 228.3 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-18031-v30.xml emd-18031-v30.xml emd-18031.xml emd-18031.xml | 16.2 KB 16.2 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
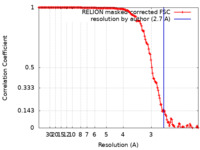
| FSC (解像度算出) |  emd_18031_fsc.xml emd_18031_fsc.xml | 14.1 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_18031.png emd_18031.png | 61 KB | ||
| Filedesc metadata |  emd-18031.cif.gz emd-18031.cif.gz | 5.8 KB | ||
| その他 |  emd_18031_half_map_1.map.gz emd_18031_half_map_1.map.gz emd_18031_half_map_2.map.gz emd_18031_half_map_2.map.gz | 193.7 MB 193.6 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18031 http://ftp.pdbj.org/pub/emdb/structures/EMD-18031 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18031 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18031 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  8pysMC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_18031.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_18031.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | The 2.7 A resolution cryo-EM reconstruction of Mycobacterium tuberculosis encapsulin containing DyP peroxidase | ||||||||||||||||||||||||||||||||||||
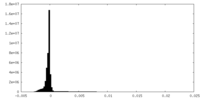
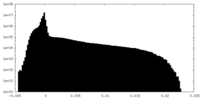
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-ハーフマップ: Half map 1
| ファイル | emd_18031_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Half map 1 | ||||||||||||
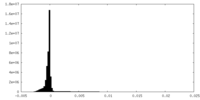
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: Half map 2
| ファイル | emd_18031_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Half map 2 | ||||||||||||
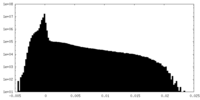
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : DyP cargo loaded encapsulin nanocompartment
| 全体 | 名称: DyP cargo loaded encapsulin nanocompartment |
|---|---|
| 要素 |
|
-超分子 #1: DyP cargo loaded encapsulin nanocompartment
| 超分子 | 名称: DyP cargo loaded encapsulin nanocompartment / タイプ: organelle_or_cellular_component / ID: 1 / 親要素: 0 / 含まれる分子: all |
|---|---|
| 由来(天然) | 生物種:  Mycobacterium tuberculosis H37Rv (結核菌) Mycobacterium tuberculosis H37Rv (結核菌) |
| 分子量 | 理論値: 1 MDa |
-分子 #1: 29 kDa antigen CFP29
| 分子 | 名称: 29 kDa antigen CFP29 / タイプ: protein_or_peptide / ID: 1 / コピー数: 60 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Mycobacterium tuberculosis H37Rv (結核菌) Mycobacterium tuberculosis H37Rv (結核菌) |
| 分子量 | 理論値: 29.554072 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MNNLYRDLAP VTEAAWAEIE LEAARTFKRH IAGRRVVDVS DPGGPVTAAV STGRLIDVKA PTNGVIAHLR ASKPLVRLRV PFTLSRNEI DDVERGSKDS DWEPVKEAAK KLAFVEDRTI FEGYSAASIE GIRSASSNPA LTLPEDPREI PDVISQALSE L RLAGVDGP ...文字列: MNNLYRDLAP VTEAAWAEIE LEAARTFKRH IAGRRVVDVS DPGGPVTAAV STGRLIDVKA PTNGVIAHLR ASKPLVRLRV PFTLSRNEI DDVERGSKDS DWEPVKEAAK KLAFVEDRTI FEGYSAASIE GIRSASSNPA LTLPEDPREI PDVISQALSE L RLAGVDGP YSVLLSADVY TKVSETSDHG YPIREHLNRL VDGDIIWAPA IDGAFVLTTR GGDFDLQLGT DVAIGYASHD TD TVRLYLQ ETLTFLCYTA EASVALSHHH HHH UniProtKB: Type 1 encapsulin shell protein |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 構成要素:
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| グリッド | モデル: Quantifoil R2/2 / 材質: COPPER / 前処理 - タイプ: GLOW DISCHARGE / 前処理 - 時間: 30 sec. | |||||||||||||||
| 凍結 | 凍結剤: NITROGEN |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 撮影したグリッド数: 1 / 実像数: 6204 / 平均電子線量: 45.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: SPOT SCAN / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 3.0 µm / 最小 デフォーカス(公称値): 0.5 µm / 倍率(公称値): 81000 |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 初期モデル | PDB ID: Chain - Chain ID: A / Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| 精密化 | プロトコル: RIGID BODY FIT / 温度因子: 26.39 当てはまり具合の基準: Cross-correlation coefficient |
| 得られたモデル |  PDB-8pys: |
 ムービー
ムービー コントローラー
コントローラー





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)