[English] 日本語
 Yorodumi
Yorodumi- EMDB-17708: Subtomogram average of Vaccinia A10 trimer with tight center from... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Subtomogram average of Vaccinia A10 trimer with tight center from in vitro cores | |||||||||
 Map data Map data | Subtomogram average of Vaccinia A10 trimer with tight center from in vitro cores | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | A10 trimer / palisade protein / tight center / VIRAL PROTEIN | |||||||||
| Biological species |  Vaccinia virus WR Vaccinia virus WR | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 8.5 Å | |||||||||
 Authors Authors | Turonova B / Liu J | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: The palisade layer of the poxvirus core is composed of flexible A10 trimers. Authors: Jiasui Liu / Simon Corroyer-Dulmont / Vojtěch Pražák / Iskander Khusainov / Karola Bahrami / Sonja Welsch / Daven Vasishtan / Agnieszka Obarska-Kosińska / Sigurdur R Thorkelsson / Kay ...Authors: Jiasui Liu / Simon Corroyer-Dulmont / Vojtěch Pražák / Iskander Khusainov / Karola Bahrami / Sonja Welsch / Daven Vasishtan / Agnieszka Obarska-Kosińska / Sigurdur R Thorkelsson / Kay Grünewald / Emmanuelle R J Quemin / Beata Turoňová / Jacomina Krijnse Locker /    Abstract: Due to its asymmetric shape, size and compactness, the structure of the infectious mature virus (MV) of vaccinia virus (VACV), the best-studied poxvirus, remains poorly understood. Instead, subviral ...Due to its asymmetric shape, size and compactness, the structure of the infectious mature virus (MV) of vaccinia virus (VACV), the best-studied poxvirus, remains poorly understood. Instead, subviral particles, in particular membrane-free viral cores, have been studied with cryo-electron microscopy. Here, we compared viral cores obtained by detergent stripping of MVs with cores in the cellular cytoplasm, early in infection. We focused on the prominent palisade layer on the core surface, combining cryo-electron tomography, subtomogram averaging and AlphaFold2 structure prediction. We showed that the palisade is composed of densely packed trimers of the major core protein A10. Trimers display a random order and their classification indicates structural flexibility. A10 on cytoplasmic cores is organized in a similar manner, indicating that the structures obtained in vitro are physiologically relevant. We discuss our results in the context of the VACV replicative cycle, and the assembly and disassembly of the infectious MV. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17708.map.gz emd_17708.map.gz | 3.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17708-v30.xml emd-17708-v30.xml emd-17708.xml emd-17708.xml | 13.6 KB 13.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17708_fsc.xml emd_17708_fsc.xml | 3.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_17708.png emd_17708.png | 85.2 KB | ||
| Filedesc metadata |  emd-17708.cif.gz emd-17708.cif.gz | 4.3 KB | ||
| Others |  emd_17708_half_map_1.map.gz emd_17708_half_map_1.map.gz emd_17708_half_map_2.map.gz emd_17708_half_map_2.map.gz | 3.1 MB 3.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17708 http://ftp.pdbj.org/pub/emdb/structures/EMD-17708 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17708 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17708 | HTTPS FTP |
-Validation report
| Summary document |  emd_17708_validation.pdf.gz emd_17708_validation.pdf.gz | 900.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_17708_full_validation.pdf.gz emd_17708_full_validation.pdf.gz | 900.3 KB | Display | |
| Data in XML |  emd_17708_validation.xml.gz emd_17708_validation.xml.gz | 9 KB | Display | |
| Data in CIF |  emd_17708_validation.cif.gz emd_17708_validation.cif.gz | 11.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17708 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17708 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17708 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17708 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_17708.map.gz / Format: CCP4 / Size: 3.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17708.map.gz / Format: CCP4 / Size: 3.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Subtomogram average of Vaccinia A10 trimer with tight center from in vitro cores | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.12 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Even half map
| File | emd_17708_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Even half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
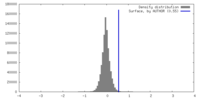
| Density Histograms |
-Half map: Odd half map
| File | emd_17708_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Odd half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
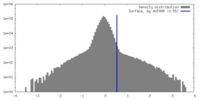
| Density Histograms |
- Sample components
Sample components
-Entire : Vaccinia virus WR
| Entire | Name:  Vaccinia virus WR Vaccinia virus WR |
|---|---|
| Components |
|
-Supramolecule #1: Vaccinia virus WR
| Supramolecule | Name: Vaccinia virus WR / type: virus / ID: 1 / Parent: 0 / NCBI-ID: 10254 / Sci species name: Vaccinia virus WR / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris X / Energy filter - Slit width: 10 eV |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Number real images: 1 / Average electron dose: 3.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)