[English] 日本語
 Yorodumi
Yorodumi- EMDB-17581: Cryo-EM structure of the c-di-GMP-bound FleQ-FleN master regulato... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the c-di-GMP-bound FleQ-FleN master regulator complex from Pseudomonas aeruginosa | |||||||||
 Map data Map data | Deep EMhancer sharpened map of the c-di-GMP-bound FleQ-FleN master regulator complex of P. aeruginosa | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Biofilm formation / c-di-GMP signaling / second messengers / bacterial secretion / gene regulation / bEBPs | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of cilium-dependent cell motility / regulation of bacterial-type flagellum-dependent cell motility / negative regulation of cell division / cyclic-di-GMP binding / positive regulation of cell-substrate adhesion / negative regulation of extracellular matrix assembly / DNA-binding transcription repressor activity / DNA-binding transcription activator activity / cis-regulatory region sequence-specific DNA binding / protein-DNA complex ...positive regulation of cilium-dependent cell motility / regulation of bacterial-type flagellum-dependent cell motility / negative regulation of cell division / cyclic-di-GMP binding / positive regulation of cell-substrate adhesion / negative regulation of extracellular matrix assembly / DNA-binding transcription repressor activity / DNA-binding transcription activator activity / cis-regulatory region sequence-specific DNA binding / protein-DNA complex / cytoplasmic side of plasma membrane / transcription cis-regulatory region binding / DNA-templated transcription / regulation of DNA-templated transcription / ATP hydrolysis activity / ATP binding / cytosol Similarity search - Function | |||||||||
| Biological species |  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) | |||||||||
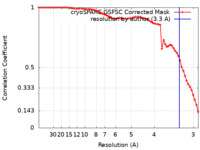
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Torres-Sanchez LT / Krasteva PV | |||||||||
| Funding support | European Union, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Structures of the FleQ-FleN master regulators reveal large-scale conformational switching in motility and biofilm control. Authors: Lucía Torres-Sánchez / Thibault Géry Sana / Marion Decossas / Yaser Hashem / Petya Violinova Krasteva /  Abstract: can cause a wide array of chronic and acute infections associated with its ability to rapidly switch between planktonic, biofilm, and dispersed lifestyles, each with a specific arsenal for bacterial ... can cause a wide array of chronic and acute infections associated with its ability to rapidly switch between planktonic, biofilm, and dispersed lifestyles, each with a specific arsenal for bacterial survival and virulence. At the cellular level, many of the physiological transitions are orchestrated by the intracellular second messenger c-di-GMP and its receptor-effector FleQ. A bacterial enhancer binding protein, FleQ acts as a master regulator of both flagellar motility and adherence factor secretion and uses remarkably different transcription activation mechanisms depending on its dinucleotide loading state, adenosine triphosphatase (ATPase) activity, interactions with polymerase sigma (σ) factors, and complexation with a second ATPase, FleN. How the FleQ-FleN tandem can exert diverse effects through recognition of a conserved FleQ binding consensus has remained enigmatic. Here, we provide cryogenic electron microscopy (cryo-EM) structures of both c-di-GMP-bound and c-di-GMP-free FleQ-FleN complexes which deepen our understanding of the proteins' (di)nucleotide-dependent conformational switching and fine-tuned roles in gene expression regulation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17581.map.gz emd_17581.map.gz | 27.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17581-v30.xml emd-17581-v30.xml emd-17581.xml emd-17581.xml | 22.8 KB 22.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17581_fsc.xml emd_17581_fsc.xml | 6.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_17581.png emd_17581.png | 80.9 KB | ||
| Filedesc metadata |  emd-17581.cif.gz emd-17581.cif.gz | 7.5 KB | ||
| Others |  emd_17581_additional_1.map.gz emd_17581_additional_1.map.gz emd_17581_half_map_1.map.gz emd_17581_half_map_1.map.gz emd_17581_half_map_2.map.gz emd_17581_half_map_2.map.gz | 15.4 MB 28.3 MB 28.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17581 http://ftp.pdbj.org/pub/emdb/structures/EMD-17581 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17581 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17581 | HTTPS FTP |
-Validation report
| Summary document |  emd_17581_validation.pdf.gz emd_17581_validation.pdf.gz | 618.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_17581_full_validation.pdf.gz emd_17581_full_validation.pdf.gz | 618.4 KB | Display | |
| Data in XML |  emd_17581_validation.xml.gz emd_17581_validation.xml.gz | 13.8 KB | Display | |
| Data in CIF |  emd_17581_validation.cif.gz emd_17581_validation.cif.gz | 17.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17581 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17581 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17581 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17581 | HTTPS FTP |
-Related structure data
| Related structure data |  8pb9MC  8p53C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17581.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17581.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Deep EMhancer sharpened map of the c-di-GMP-bound FleQ-FleN master regulator complex of P. aeruginosa | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.419 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Unsharpened map of the c-di-GMP-bound FleQ-FleN master regulator...
| File | emd_17581_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map of the c-di-GMP-bound FleQ-FleN master regulator complex of P. aeruginosa | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half-map of the c-di-GMP-bound FleQ-FleN master regulator complex...
| File | emd_17581_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map of the c-di-GMP-bound FleQ-FleN master regulator complex of P. aeruginosa | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half-map of the c-di-GMP-bound FleQ-FleN master regulator complex...
| File | emd_17581_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map of the c-di-GMP-bound FleQ-FleN master regulator complex of P. aeruginosa | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : C-di-GMP bound FleQ-FleN complex from Pseudomonas aeruginosa
| Entire | Name: C-di-GMP bound FleQ-FleN complex from Pseudomonas aeruginosa |
|---|---|
| Components |
|
-Supramolecule #1: C-di-GMP bound FleQ-FleN complex from Pseudomonas aeruginosa
| Supramolecule | Name: C-di-GMP bound FleQ-FleN complex from Pseudomonas aeruginosa type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 Details: C-di-GMP bound complex between FleQ REC and AAA+ domains and FleN D48A mutant from Pseudomonas aeruginosa |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) |
| Molecular weight | Theoretical: 233 KDa |
-Macromolecule #1: Transcriptional regulator FleQ
| Macromolecule | Name: Transcriptional regulator FleQ / type: protein_or_peptide / ID: 1 Details: FleQ-receiver and AAA+ domains of Pseudomonas aeruginosa FleQ Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) |
| Molecular weight | Theoretical: 44.884316 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSWRETKLL LIDDNLDRSR DLAVILNFLG EDQLTCNSED WREVAAGLSN SREALCVLLG SVESKGGAVE LLKQLASWDE YLPILLIGE PAPADWPEEL RRRVLASLEM PPSYNKLLDS LHRAQVYREM YDQARERGRS REPNLFRSLV GTSRAIQQVR Q MMQQVADT ...String: MGSWRETKLL LIDDNLDRSR DLAVILNFLG EDQLTCNSED WREVAAGLSN SREALCVLLG SVESKGGAVE LLKQLASWDE YLPILLIGE PAPADWPEEL RRRVLASLEM PPSYNKLLDS LHRAQVYREM YDQARERGRS REPNLFRSLV GTSRAIQQVR Q MMQQVADT DASVLILGES GTGKEVVARN LHYHSKRREG PFVPVNCGAI PAELLESELF GHEKGAFTGA ITSRAGRFEL AN GGTLFLD EIGDMPLPMQ VKLLRVLQER TFERVGSNKT QNVDVRIIAA THKNLEKMIE DGTFREDLYY RLNVFPIEMA PLR ERVEDI ALLLNELISR MEHEKRGSIR FNSAAIMSLC RHDWPGNVRE LANLVERLAI MHPYGVIGVG ELPKKFRHVD UniProtKB: Transcriptional regulator FleQ |
-Macromolecule #2: Antiactivator FleN
| Macromolecule | Name: Antiactivator FleN / type: protein_or_peptide / ID: 2 / Details: Pseudomonas aeruginosa FleN D48A mutant / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) |
| Molecular weight | Theoretical: 30.327141 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPMGSKQMGS MHPVQVIAVT GGKGGVGKTN VSVNLALALA DLGRRVMLLD AALGLANVDV LLGLTPKRTL ADVIEGRCEL RDVLLLGPG GVRIVPAASG TQSMVHLSPM QHAGLIQAFS DISDNLDVLV VDTAAGIGDS VVSFVRAAQE VLLVVCDEPT S ITDAYALI ...String: GPMGSKQMGS MHPVQVIAVT GGKGGVGKTN VSVNLALALA DLGRRVMLLD AALGLANVDV LLGLTPKRTL ADVIEGRCEL RDVLLLGPG GVRIVPAASG TQSMVHLSPM QHAGLIQAFS DISDNLDVLV VDTAAGIGDS VVSFVRAAQE VLLVVCDEPT S ITDAYALI KLLNRDHGMT RFRVLANMAH SPQEGRNLFA KLTKVTDRFL DVALQYVGVI PYDESVRKAV QKQRAVYEAF PR SKASLAF KAVAQKVDSW PLPANPRGHL EFFVERLVQH PATGSAV UniProtKB: Antiactivator FleN |
-Macromolecule #3: 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydr...
| Macromolecule | Name: 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one) type: ligand / ID: 3 / Number of copies: 2 / Formula: C2E |
|---|---|
| Molecular weight | Theoretical: 690.411 Da |
| Chemical component information | 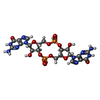 ChemComp-C2E: |
-Macromolecule #4: PHOSPHOMETHYLPHOSPHONIC ACID ADENYLATE ESTER
| Macromolecule | Name: PHOSPHOMETHYLPHOSPHONIC ACID ADENYLATE ESTER / type: ligand / ID: 4 / Number of copies: 2 / Formula: ACP |
|---|---|
| Molecular weight | Theoretical: 505.208 Da |
| Chemical component information |  ChemComp-ACP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Details: 20 mM HEPES pH 8.0, 250mM NaCl, 2mM MgCl2, and 2% glycerol, 4 uM c-di-GMP, 200 uM ACP |
|---|---|
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: triple deposition. |
| Details | FleN-D48A and FleQ (REC-AAA+) coexpression from the pProEx-Htb and pRSFDuet1 vectors. Purified via the HRV3c- cleavable N-terminal hexahistidine tag on FleN |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 1 / Number real images: 14640 / Average electron dose: 50.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: Other / Chain - Initial model type: experimental model Details: the model was interpreted by modular fitting of partial crystal structures and refined against the electron density |
|---|---|
| Details | Crystal structures fitting Refined in Phenix, coot and Namdinator |
| Refinement | Space: REAL |
| Output model |  PDB-8pb9: |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X