+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | S.cerevisiae THO complex from endogenous nuclear mRNPs | ||||||||||||
 Map data Map data | dimer masked map | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Complex / Nuclear / Dimer / NUCLEAR PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationnucleoplasmic THO complex / cellular response to azide / THO complex / THO complex part of transcription export complex / positive regulation of transcription elongation by RNA polymerase I / transcription export complex / Cdc73/Paf1 complex / mRNA 3'-end processing / positive regulation of transcription by RNA polymerase I / transcription-coupled nucleotide-excision repair ...nucleoplasmic THO complex / cellular response to azide / THO complex / THO complex part of transcription export complex / positive regulation of transcription elongation by RNA polymerase I / transcription export complex / Cdc73/Paf1 complex / mRNA 3'-end processing / positive regulation of transcription by RNA polymerase I / transcription-coupled nucleotide-excision repair / mRNA export from nucleus / stress granule assembly / transcription elongation by RNA polymerase II / mRNA processing / DNA recombination / nucleic acid binding / chromosome, telomeric region / molecular adaptor activity / mRNA binding / nucleus Similarity search - Function | ||||||||||||
| Biological species |   | ||||||||||||
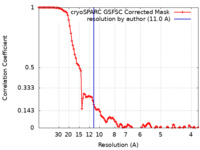
| Method | single particle reconstruction / cryo EM / Resolution: 11.0 Å | ||||||||||||
 Authors Authors | Bonneau F / Schaefer IB / Conti E | ||||||||||||
| Funding support | European Union,  Germany, Germany,  Denmark, 3 items Denmark, 3 items
| ||||||||||||
 Citation Citation |  Journal: Genes Dev / Year: 2023 Journal: Genes Dev / Year: 2023Title: Nuclear mRNPs are compact particles packaged with a network of proteins promoting RNA-RNA interactions. Authors: Fabien Bonneau / Jérôme Basquin / Barbara Steigenberger / Tillman Schäfer / Ingmar B Schäfer / Elena Conti /  Abstract: Messenger RNAs (mRNAs) are at the center of the central dogma of molecular biology. In eukaryotic cells, these long ribonucleic acid polymers do not exist as naked transcripts; rather, they associate ...Messenger RNAs (mRNAs) are at the center of the central dogma of molecular biology. In eukaryotic cells, these long ribonucleic acid polymers do not exist as naked transcripts; rather, they associate with mRNA-binding proteins to form messenger ribonucleoprotein (mRNP) complexes. Recently, global proteomic and transcriptomic studies have provided comprehensive inventories of mRNP components. However, knowledge of the molecular features of distinct mRNP populations has remained elusive. We purified endogenous nuclear mRNPs from by harnessing the mRNP biogenesis factors THO and Sub2 in biochemical procedures optimized to preserve the integrity of these transient ribonucleoprotein assemblies. We found that these mRNPs are compact particles that contain multiple copies of Yra1, an essential protein with RNA-annealing properties. To investigate their molecular and architectural organization, we used a combination of proteomics, RNA sequencing, cryo-electron microscopy, cross-linking mass spectrometry, structural models, and biochemical assays. Our findings indicate that yeast nuclear mRNPs are packaged around an intricate network of interconnected proteins capable of promoting RNA-RNA interactions via their positively charged intrinsically disordered regions. The evolutionary conservation of the major mRNA-packaging factor (yeast Yra1 and Aly/REF in metazoans) points toward a general paradigm governing nuclear mRNP packaging. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16841.map.gz emd_16841.map.gz | 121.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16841-v30.xml emd-16841-v30.xml emd-16841.xml emd-16841.xml | 22.9 KB 22.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16841_fsc.xml emd_16841_fsc.xml | 10.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_16841.png emd_16841.png | 27.4 KB | ||
| Masks |  emd_16841_msk_1.map emd_16841_msk_1.map | 129.7 MB |  Mask map Mask map | |
| Others |  emd_16841_half_map_1.map.gz emd_16841_half_map_1.map.gz emd_16841_half_map_2.map.gz emd_16841_half_map_2.map.gz | 120.3 MB 120.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16841 http://ftp.pdbj.org/pub/emdb/structures/EMD-16841 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16841 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16841 | HTTPS FTP |
-Validation report
| Summary document |  emd_16841_validation.pdf.gz emd_16841_validation.pdf.gz | 712.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_16841_full_validation.pdf.gz emd_16841_full_validation.pdf.gz | 712.5 KB | Display | |
| Data in XML |  emd_16841_validation.xml.gz emd_16841_validation.xml.gz | 19.3 KB | Display | |
| Data in CIF |  emd_16841_validation.cif.gz emd_16841_validation.cif.gz | 25.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16841 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16841 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16841 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16841 | HTTPS FTP |
-Related structure data
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16841.map.gz / Format: CCP4 / Size: 129.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16841.map.gz / Format: CCP4 / Size: 129.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | dimer masked map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.885 Å | ||||||||||||||||||||||||||||||||||||
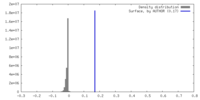
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16841_msk_1.map emd_16841_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
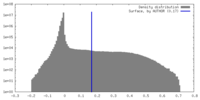
| Density Histograms |
-Half map: dimer half map A
| File | emd_16841_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | dimer half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
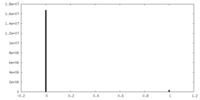
| Density Histograms |
-Half map: dimer half map B
| File | emd_16841_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | dimer half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
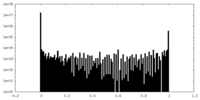
| Density Histograms |
- Sample components
Sample components
-Entire : THO complex dimer
| Entire | Name: THO complex dimer |
|---|---|
| Components |
|
-Supramolecule #1: THO complex dimer
| Supramolecule | Name: THO complex dimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: From nuclear mRNPs isolated by bi-molecular affinity purification using Sub2 and Hpr1 as baits, then treated with Benzonase. |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: THO complex subunit 2
| Macromolecule | Name: THO complex subunit 2 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MAEQTLLSKL NALSQKVIPP ASPSQASILT EEVIRNWPER SKTLCSDFTA LESNDEKEDW LRTLFIELFD FINKNDENSP LKLSDVASFT NELVNHERQV SQASIVGKMF IAVSSTVPNI NDLTTISLCK LIPSLHEELF KFSWISSKLL NKEQTTLLRH LLKKSKYELK ...String: MAEQTLLSKL NALSQKVIPP ASPSQASILT EEVIRNWPER SKTLCSDFTA LESNDEKEDW LRTLFIELFD FINKNDENSP LKLSDVASFT NELVNHERQV SQASIVGKMF IAVSSTVPNI NDLTTISLCK LIPSLHEELF KFSWISSKLL NKEQTTLLRH LLKKSKYELK KYNLLVENSV GYGQLVALLI LAYYDPDNFS KVSAYLKEIY HIMGKYSLDS IRTLDVILNV SSQFITEGYK FFIALLRKSD SWPSSHVANN SNYSSLNEGG NMIAANIISF NLSQYNEEVD KENYERYMDM CCILLKNGFV NFYSIWDNVK PEMEFLQEYI QNLETELEEE STKGVENPLA MAAALSTENE TDEDNALVVN DDVNMKDKIS EETNADIESK GKQKTQQDIL LFGKIKLLER LLIHGCVIPV IHVLKQYPKV LYVSESLSRY LGRVFEYLLN PLYTSMTSSG ESKDMATALM ITRIDNGILA HKPRLIHKYK THEPFESLEL NSSYVFYYSE WNSNLTPFAS VNDLFENSHI YLSIIGPYLG RIPTLLSKIS RIGVADIQKN HGSESLHVTI DKWIDYVRKF IFPATSLLQN NPIATSEVYE LMKFFPFEKR YFIYNEMMTK LSQDILPLKV SFNKAEREAK SILKALSIDT IAKESRRFAK LISTNPLASL VPAVKQIENY DKVSELVVYT TKYFNDFAYD VLQFVLLLRL TYNRPAVQFD GVNQAMWVQR LSIFIAGLAK NCPNMDISNI ITYILKTLHN GNIIAVSILK ELIITVGGIR DLNEVNMKQL LMLNSGSPLK QYARHLIYDF RDDNSVISSR LTSFFTDQSA ISEIILLLYT LNLKANTQNS HYKILSTRCD EMNTLLWSFI ELIKHCLKGK AFEENVLPFV ELNNRFHLST PWTFHIWRDY LDNQLNSNEN FSIDELIEGA EFSDVDLTKI SKDLFTTFWR LSLYDIHFDK SLYDERKNAL SGENTGHMSN RKKHLIQNQI KDILVTGISH QRAFKKTSEF ISEKSNVWNK DCGEDQIKIF LQNCVVPRVL FSPSDALFSS FFIFMAFRTE NLMSILNTCI TSNILKTLLF CCTSSEAGNL GLFFTDVLKK LEKMRLNGDF NDQASRKLYE WHSVITEQVI DLLSEKNYMS IRNGIEFMKH VTSVFPVVKA HIQLVYTTLE ENLINEERED IKLPSSALIG HLKARLKDAL ELDEFCTLTE EEAEQKRIRE MELEEIKNYE TACQNEQKQV ALRKQLELNK SQRLQNDPPK SVASGSAGLN SKDRYTYSRN EPVIPTKPSS SQWSYSKVTR HVDDINHYLA TNHLQKAISL VENDDETRNL RKLSKQNMPI FDFRNSTLEI FERYFRTLIQ NPQNPDFAEK IDSLKRYIKN ISREPYPDTT SSYSEAAAPE YTKRSSRYSG NAGGKDGYGS SNYRGPSNDR SAPKNIKPIS SYAHKRSELP TRPSKSKTYN DRSRALRPTG PDRGDGFDQR DNRLREEYKK NSSQRSQLRF PEKPFQEGKD SSKANPYQAS SYKRDSPSEN EEKPNKRFKK DETIRNKFQT QDYRNTRDSG AAHRANENQR YNGNRKSNTQ ALPQGPKGGN YVSRYQR UniProtKB: THO complex subunit 2 |
-Macromolecule #2: THO complex subunit HPR1
| Macromolecule | Name: THO complex subunit HPR1 / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MSNTEELIQN SIGFLQKTFK ALPVSFDSIR HEPLPSSMLH ASVLNFEWEP LEKNISAIHD RDSLIDIILK RFIIDSMTNA IEDEEENNLE KGLLNSCIGL DFVYNSRFNR SNPASWGNTF FELFSTIIDL LNSPSTFLKF WPYAESRIEW FKMNTSVEPV SLGESNLISY ...String: MSNTEELIQN SIGFLQKTFK ALPVSFDSIR HEPLPSSMLH ASVLNFEWEP LEKNISAIHD RDSLIDIILK RFIIDSMTNA IEDEEENNLE KGLLNSCIGL DFVYNSRFNR SNPASWGNTF FELFSTIIDL LNSPSTFLKF WPYAESRIEW FKMNTSVEPV SLGESNLISY KQPLYEKLRH WNDILAKLEN NDILNTVKHY NMKYKLENFL SELLPINEES NFNRSASISA LQESDNEWNR SARERESNRS SDVIFAADYN FVFYHLIICP IEFAFSDLEY KNDVDRSLSP LLDAILEIEE NFYSKIKMNN RTRYSLEEAL NTEYYANYDV MTPKLPVYMK HSNAMKMDRN EFWANLQNIK ESDDYTLRPT IMDISLSNTT CLYKQLTQED DDYYRKQFIL QLCFTTNLIR NLISSDETRN FYKSCYLREN PLSDIDFENL DEVNKKRGLN LCSYICDNRV LKFYKIKDPD FYRVIRKLMS SDEKFTTAKI DGFKEFQNFR ISKEKIPPPA FDETFKKFTF IKMGNKLINN VWKIPTGLDK IEQEVKKPEG VYEAAQAKWE SKISSETSGG EAKDEIIRQW QTLRFLRSRY LFDFDKVNEK TGVDGLFEEP RKVEALDDSF KEKLLYKINQ EHRKKLQDAR EYKIGKERKK RALEEEASFP EREQKIKSQR INSASQTEGD ELKSEQTQPK GEISEENTKI KSSEVSSQDP DSGVAGEFAP QNTTAQLENP KTEDNNAATS NISNGSSTQD MKGSGSGSGS GSSAWSHPQF EKGGGSGGGS GGSAWSHPQF EK UniProtKB: THO complex subunit HPR1 |
-Macromolecule #3: THO complex subunit MFT1
| Macromolecule | Name: THO complex subunit MFT1 / type: protein_or_peptide / ID: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MPLSQKQIDQ VRTKVHYSEV DTPFNKYLDI LGKVTKLTGS IINGTLSNDD SKIEKLTEQN ISQLKESAHL RFLDLQSSID TKKVADENWE TCQQETLAKL ENLKDKLPDI KSIHSKLLLR IGKLQGLYDS VQVINREVEG LSEGRTSLVV TRAEWEKELG TDLVKFLIEK ...String: MPLSQKQIDQ VRTKVHYSEV DTPFNKYLDI LGKVTKLTGS IINGTLSNDD SKIEKLTEQN ISQLKESAHL RFLDLQSSID TKKVADENWE TCQQETLAKL ENLKDKLPDI KSIHSKLLLR IGKLQGLYDS VQVINREVEG LSEGRTSLVV TRAEWEKELG TDLVKFLIEK NYLKLVDPGL KKDSSEERYR IYDDFSKGPK ELESINASMK SDIENVRQEV SSYKEKWLRD AEIFGKITSI FKEELLKRDG LLNEAEGDNI DEDYESDEDE ERKERFKRQR SMVEVNTIEN VDEKEESDHE YDDQEDEENE EEDDMEVDVE DIKEDNEVDG ESSQQEDNSR QGNNEETDKE TGVIEEPDAV NDAEEADSDH SSRKLGGTTS DFSASSSVEE VK UniProtKB: THO complex subunit MFT1 |
-Macromolecule #4: Protein TEX1
| Macromolecule | Name: Protein TEX1 / type: protein_or_peptide / ID: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MSTIGAVDIL NQKTITSEVA ASVTSKYLQS TFSKGNTSHI EDKRFIHVSS RSHSRFTSTP ITPNEILSLK FHVSGSSMAY SRMDGSLTVW FIKDASFDKS VEVYIPDCCG SDKLATDLSW NPTSLNQIAV VSNSSEISLL LINEKSLTAS KLRTLSLGSK TKVNTCLYDP ...String: MSTIGAVDIL NQKTITSEVA ASVTSKYLQS TFSKGNTSHI EDKRFIHVSS RSHSRFTSTP ITPNEILSLK FHVSGSSMAY SRMDGSLTVW FIKDASFDKS VEVYIPDCCG SDKLATDLSW NPTSLNQIAV VSNSSEISLL LINEKSLTAS KLRTLSLGSK TKVNTCLYDP LGNWLLAATK SEKIYLFDVK KDHSSVCSLN ISDISQEDND VVYSLAWSNG GSHIFIGFKS GYLAILKAKH GILEVCTKIK AHTGPITEIK MDPWGRNFIT GSIDGNCYVW NMKSLCCELI INDLNSAVTT LDVCHLGKIL GICTEDEMVY FYDLNSGNLL HSKSLANYKT DPVLKFYPDK SWYIMSGKND TLSNHFVKNE KNLITYWKDM FDNTMIEKRR KNNGGGNNHN KRTSKNTDRI GKDRPSRFNS KK UniProtKB: Protein TEX1 |
-Macromolecule #5: THO complex subunit THP2
| Macromolecule | Name: THO complex subunit THP2 / type: protein_or_peptide / ID: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MTKEEGRTYF ESLCEEEQSL QESQTHLLNI LDILSVLADP RSSDDLLTES LKKLPDLHRE LINSSIRLRY DKYQTREAQL LEDTKTGRDV AAGVQNPKSI SEYYSTFEHL NRDTLRYINL LKRLSVDLAK QVEVSDPSVT VYEMDKWVPS EKLQGILEQY CAPDTDIRGV ...String: MTKEEGRTYF ESLCEEEQSL QESQTHLLNI LDILSVLADP RSSDDLLTES LKKLPDLHRE LINSSIRLRY DKYQTREAQL LEDTKTGRDV AAGVQNPKSI SEYYSTFEHL NRDTLRYINL LKRLSVDLAK QVEVSDPSVT VYEMDKWVPS EKLQGILEQY CAPDTDIRGV DAQIKNYLDQ IKMARAKFGL ENKYSLKERL STLTKELNHW RKEWDDIEML MFGDDAHSMK KMIQKIDSLK SEINAPSESY PVDKEGDIVL E UniProtKB: THO complex subunit THP2 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. / Pretreatment - Atmosphere: AIR | ||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-40 / Number grids imaged: 1 / Number real images: 2258 / Average exposure time: 16.0 sec. / Average electron dose: 59.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.62 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 22000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)