+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of TolQR complex from E.coli | |||||||||
 Map data Map data | One of maps used in refinement, Z-flipped for model building. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TolQ / TolR / Tol-Pal / complex / inner membrane / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of membrane invagination / cell septum assembly / bacteriocin transport / protein import / cell envelope / cell division site / transmembrane transporter activity / protein transport / cell division / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   | |||||||||
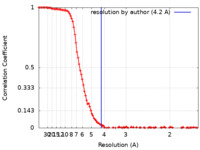
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | |||||||||
 Authors Authors | Webby MN / Kleanthous C / Press CE | |||||||||
| Funding support |  United Kingdom, European Union, 2 items United Kingdom, European Union, 2 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Tunable force transduction through the cell envelope. Authors: Daniel P Williams-Jones / Melissa N Webby / Cara E Press / Jan M Gradon / Sophie R Armstrong / Joanna Szczepaniak / Colin Kleanthous /  Abstract: The outer membrane (OM) of Gram-negative bacteria is not energised and so processes requiring a driving force must connect to energy-transduction systems in the inner membrane (IM). Tol (Tol-Pal) and ...The outer membrane (OM) of Gram-negative bacteria is not energised and so processes requiring a driving force must connect to energy-transduction systems in the inner membrane (IM). Tol (Tol-Pal) and Ton are related, proton motive force- (PMF-) coupled assemblies that stabilise the OM and import essential nutrients, respectively. Both rely on proton-harvesting IM motor (stator) complexes, which are homologues of the flagellar stator unit Mot, to transduce force to the OM through elongated IM force transducer proteins, TolA and TonB, respectively. How PMF-driven motors in the IM generate mechanical work at the OM via force transducers is unknown. Here, using cryoelectron microscopy, we report the 4.3Å structure of the TolQR motor complex. The structure reaffirms the 5:2 stoichiometry seen in Ton and Mot and, with motor subunits related to each other by 10 to 16° rotation, supports rotary motion as the default for these complexes. We probed the mechanism of force transduction to the OM through in vivo assays of chimeric TolA/TonB proteins where sections of their structurally divergent, periplasm-spanning domains were swapped or replaced by an intrinsically disordered sequence. We find that TolA mutants exhibit a spectrum of force output, which is reflected in their respective abilities to both stabilise the OM and import cytotoxic colicins across the OM. Our studies demonstrate that structural rigidity of force transducer proteins, rather than any particular structural form, drives the efficient conversion of PMF-driven rotary motions of 5:2 motor complexes into physiologically relevant force at the OM. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16816.map.gz emd_16816.map.gz | 96.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16816-v30.xml emd-16816-v30.xml emd-16816.xml emd-16816.xml | 18.5 KB 18.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16816_fsc.xml emd_16816_fsc.xml | 13.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_16816.png emd_16816.png | 51.3 KB | ||
| Filedesc metadata |  emd-16816.cif.gz emd-16816.cif.gz | 5.9 KB | ||
| Others |  emd_16816_additional_1.map.gz emd_16816_additional_1.map.gz emd_16816_half_map_1.map.gz emd_16816_half_map_1.map.gz emd_16816_half_map_2.map.gz emd_16816_half_map_2.map.gz | 96.9 MB 95.5 MB 95.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16816 http://ftp.pdbj.org/pub/emdb/structures/EMD-16816 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16816 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16816 | HTTPS FTP |
-Related structure data
| Related structure data |  8odtMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16816.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16816.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | One of maps used in refinement, Z-flipped for model building. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.832 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Class one, from 3D classification.
| File | emd_16816_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Class one, from 3D classification. | ||||||||||||
| Projections & Slices |
| ||||||||||||
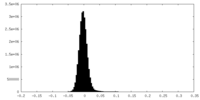
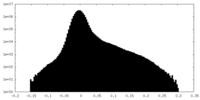
| Density Histograms |
-Half map: Half map A
| File | emd_16816_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A
| File | emd_16816_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : TolQ-TolR 5:2 complex
| Entire | Name: TolQ-TolR 5:2 complex |
|---|---|
| Components |
|
-Supramolecule #1: TolQ-TolR 5:2 complex
| Supramolecule | Name: TolQ-TolR 5:2 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 170 KDa |
-Macromolecule #1: Tol-Pal system protein TolQ
| Macromolecule | Name: Tol-Pal system protein TolQ / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 25.623662 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTDMNILDLF LKASLLVKLI MLILIGFSIA SWAIIIQRTR ILNAAAREAE AFEDKFWSGI ELSRLYQESQ GKRDNLTGSE QIFYSGFKE FVRLHRANSH APEAVVEGAS RAMRISMNRE LENLETHIPF LGTVGSISPY IGLFGTVWGI MHAFIALGAV K QATLQMVA ...String: MTDMNILDLF LKASLLVKLI MLILIGFSIA SWAIIIQRTR ILNAAAREAE AFEDKFWSGI ELSRLYQESQ GKRDNLTGSE QIFYSGFKE FVRLHRANSH APEAVVEGAS RAMRISMNRE LENLETHIPF LGTVGSISPY IGLFGTVWGI MHAFIALGAV K QATLQMVA PGIAEALIAT AIGLFAAIPA VMAYNRLNQR VNKLELNYDN FMEEFTAILH RQAFTVSESN KG UniProtKB: Tol-Pal system protein TolQ |
-Macromolecule #2: Tol-Pal system protein TolR
| Macromolecule | Name: Tol-Pal system protein TolR / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 20.600367 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MARARGRGRR DLKSEINIVP LLDVLLVLLL IFMATAPIIT QSVEVDLPDA TESQAVSSND NPPVIVEVSG IGQYTVVVEK DRLERLPPE QVVAEVSSRF KANPKTVFLI GGAKDVPYDE IIKALNLLHS AGVKSVGLMT QPILEENLYF QGQFGSWSHP Q FEKGGGSG GGSGGGSWSH PQFEKHHHHH H UniProtKB: Tol-Pal system protein TolR |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 7.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 15.0 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 281 K / Instrument: FEI VITROBOT MARK IV / Details: Blot force 10 Blot time 3 sec Wait time 2 sec. |
| Details | sample purified by affinity chromatography (His-tagged TolR) and SEC before application to grids in 50 mM Tris/HCl pH 7.5, 300 mM NaCl, 2 mM EDTA, 0.01% LMNG |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 46.39 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)