+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Wild type ATTR amyloid fibril from senile systemic amyloidosis | ||||||||||||
 マップデータ マップデータ | |||||||||||||
 試料 試料 |
| ||||||||||||
 キーワード キーワード | Transthyretin amyliod fibril / PROTEIN FIBRIL | ||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報Defective visual phototransduction due to STRA6 loss of function / negative regulation of glomerular filtration / The canonical retinoid cycle in rods (twilight vision) / purine nucleobase metabolic process / hormone binding / Non-integrin membrane-ECM interactions / phototransduction, visible light / molecular sequestering activity / retinoid metabolic process / Retinoid metabolism and transport ...Defective visual phototransduction due to STRA6 loss of function / negative regulation of glomerular filtration / The canonical retinoid cycle in rods (twilight vision) / purine nucleobase metabolic process / hormone binding / Non-integrin membrane-ECM interactions / phototransduction, visible light / molecular sequestering activity / retinoid metabolic process / Retinoid metabolism and transport / hormone activity / azurophil granule lumen / Amyloid fiber formation / Neutrophil degranulation / protein-containing complex binding / protein-containing complex / : / extracellular exosome / extracellular region / identical protein binding 類似検索 - 分子機能 | ||||||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | ||||||||||||
| 手法 | らせん対称体再構成法 / クライオ電子顕微鏡法 / 解像度: 2.78 Å | ||||||||||||
 データ登録者 データ登録者 | Schmidt M / Steinebrei M | ||||||||||||
| 資金援助 |  ドイツ, 1件 ドイツ, 1件
| ||||||||||||
 引用 引用 |  ジャーナル: Nat Commun / 年: 2022 ジャーナル: Nat Commun / 年: 2022タイトル: Cryo-EM structure of an ATTRwt amyloid fibril from systemic non-hereditary transthyretin amyloidosis. 著者: Maximilian Steinebrei / Juliane Gottwald / Julian Baur / Christoph Röcken / Ute Hegenbart / Stefan Schönland / Matthias Schmidt /  要旨: Wild type transthyretin-derived amyloid (ATTRwt) is the major component of non-hereditary transthyretin amyloidosis. Its accumulation in the heart of elderly patients is life threatening. A variety ...Wild type transthyretin-derived amyloid (ATTRwt) is the major component of non-hereditary transthyretin amyloidosis. Its accumulation in the heart of elderly patients is life threatening. A variety of genetic variants of transthyretin can lead to hereditary transthyretin amyloidosis, which shows different clinical symptoms, like age of onset and pattern of organ involvement. However, in the case of non-hereditary transthyretin amyloidosis ATTRwt fibril deposits are located primarily in heart tissue. In this structural study we analyzed ATTRwt amyloid fibrils from the heart of a patient with non-hereditary transthyretin amyloidosis. We present a 2.78 Å reconstructed density map of these ATTRwt fibrils using cryo electron microscopy and compare it with previously published V30M variants of ATTR fibrils extracted from heart and eye of different patients. All structures show a remarkably similar spearhead like shape in their cross section, formed by the same N- and C-terminal fragments of transthyretin with some minor differences. This demonstrates common features for ATTR fibrils despite differences in mutations and patients. | ||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_15361.map.gz emd_15361.map.gz | 495.2 KB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-15361-v30.xml emd-15361-v30.xml emd-15361.xml emd-15361.xml | 13.4 KB 13.4 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
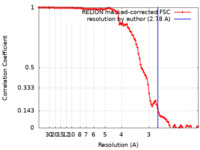
| FSC (解像度算出) |  emd_15361_fsc.xml emd_15361_fsc.xml | 10.6 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_15361.png emd_15361.png | 8.3 MB | ||
| Filedesc metadata |  emd-15361.cif.gz emd-15361.cif.gz | 4.8 KB | ||
| その他 |  emd_15361_half_map_1.map.gz emd_15361_half_map_1.map.gz emd_15361_half_map_2.map.gz emd_15361_half_map_2.map.gz | 80.6 MB 80.6 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15361 http://ftp.pdbj.org/pub/emdb/structures/EMD-15361 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15361 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15361 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  8adeMC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_15361.map.gz / 形式: CCP4 / 大きさ: 103 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_15361.map.gz / 形式: CCP4 / 大きさ: 103 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||
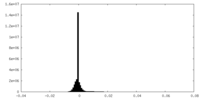
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-ハーフマップ: #1
| ファイル | emd_15361_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
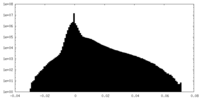
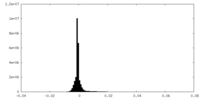
| 密度ヒストグラム |
-ハーフマップ: #2
| ファイル | emd_15361_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
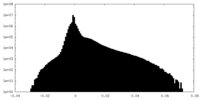
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : Transthyretin amyolid fibrils
| 全体 | 名称: Transthyretin amyolid fibrils |
|---|---|
| 要素 |
|
-超分子 #1: Transthyretin amyolid fibrils
| 超分子 | 名称: Transthyretin amyolid fibrils / タイプ: tissue / ID: 1 / 親要素: 0 / 含まれる分子: all |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) / 器官: Heart / 組織: Heart muscle Homo sapiens (ヒト) / 器官: Heart / 組織: Heart muscle |
-分子 #1: Transthyretin
| 分子 | 名称: Transthyretin / タイプ: protein_or_peptide / ID: 1 / コピー数: 7 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 13.77736 KDa |
| 配列 | 文字列: GPTGTGESKC PLMVKVLDAV RGSPAINVAV HVFRKAADDT WEPFASGKTS ESGELHGLTT EEEFVEGIYK VEIDTKSYWK ALGISPFHE HAEVVFTAND SGPRRYTIAA LLSPYSYSTT AVVTNPKE UniProtKB: Transthyretin |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | らせん対称体再構成法 |
| 試料の集合状態 | helical array |
- 試料調製
試料調製
| 緩衝液 | pH: 7 |
|---|---|
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 85 % / チャンバー内温度: 295.1500 K / 装置: LEICA EM GP |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 QUANTUM (4k x 4k) 検出モード: COUNTING / 平均電子線量: 44.9 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: SPOT SCAN / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2.5 µm / 最小 デフォーカス(公称値): 1.2 µm |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)