[English] 日本語
 Yorodumi
Yorodumi- EMDB-15132: Tomogram of a late endosome of A549-IFITM3 cells infected with in... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 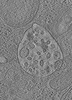 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure S8) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | IFITM3 / influenza A virus / membrane fusion / IMMUNE SYSTEM | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | electron tomography / cryo EM | |||||||||
 Authors Authors | Klein S / Chlanda P | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Host Microbe / Year: 2023 Journal: Cell Host Microbe / Year: 2023Title: IFITM3 blocks influenza virus entry by sorting lipids and stabilizing hemifusion. Authors: Steffen Klein / Gonen Golani / Fabio Lolicato / Carmen Lahr / Daniel Beyer / Alexia Herrmann / Moritz Wachsmuth-Melm / Nina Reddmann / Romy Brecht / Mehdi Hosseinzadeh / Androniki Kolovou / ...Authors: Steffen Klein / Gonen Golani / Fabio Lolicato / Carmen Lahr / Daniel Beyer / Alexia Herrmann / Moritz Wachsmuth-Melm / Nina Reddmann / Romy Brecht / Mehdi Hosseinzadeh / Androniki Kolovou / Jana Makroczyova / Sarah Peterl / Martin Schorb / Yannick Schwab / Britta Brügger / Walter Nickel / Ulrich S Schwarz / Petr Chlanda /   Abstract: Interferon-induced transmembrane protein 3 (IFITM3) inhibits the entry of numerous viruses through undefined molecular mechanisms. IFITM3 localizes in the endosomal-lysosomal system and specifically ...Interferon-induced transmembrane protein 3 (IFITM3) inhibits the entry of numerous viruses through undefined molecular mechanisms. IFITM3 localizes in the endosomal-lysosomal system and specifically affects virus fusion with target cell membranes. We found that IFITM3 induces local lipid sorting, resulting in an increased concentration of lipids disfavoring viral fusion at the hemifusion site. This increases the energy barrier for fusion pore formation and the hemifusion dwell time, promoting viral degradation in lysosomes. In situ cryo-electron tomography captured IFITM3-mediated arrest of influenza A virus membrane fusion. Observation of hemifusion diaphragms between viral particles and late endosomal membranes confirmed hemifusion stabilization as a molecular mechanism of IFITM3. The presence of the influenza fusion protein hemagglutinin in post-fusion conformation close to hemifusion sites further indicated that IFITM3 does not interfere with the viral fusion machinery. Collectively, these findings show that IFITM3 induces lipid sorting to stabilize hemifusion and prevent virus entry into target cells. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15132.map.gz emd_15132.map.gz | 5.4 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15132-v30.xml emd-15132-v30.xml emd-15132.xml emd-15132.xml | 9.7 KB 9.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_15132.png emd_15132.png | 174 KB | ||
| Filedesc metadata |  emd-15132.cif.gz emd-15132.cif.gz | 3.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15132 http://ftp.pdbj.org/pub/emdb/structures/EMD-15132 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15132 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15132 | HTTPS FTP |
-Validation report
| Summary document |  emd_15132_validation.pdf.gz emd_15132_validation.pdf.gz | 434.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_15132_full_validation.pdf.gz emd_15132_full_validation.pdf.gz | 434 KB | Display | |
| Data in XML |  emd_15132_validation.xml.gz emd_15132_validation.xml.gz | 5 KB | Display | |
| Data in CIF |  emd_15132_validation.cif.gz emd_15132_validation.cif.gz | 6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15132 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15132 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15132 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15132 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15132.map.gz / Format: CCP4 / Size: 9.3 GB / Type: IMAGE STORED AS SIGNED BYTE Download / File: emd_15132.map.gz / Format: CCP4 / Size: 9.3 GB / Type: IMAGE STORED AS SIGNED BYTE | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 2.671 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : A549 cell stably overexpressing IFITM3
| Entire | Name: A549 cell stably overexpressing IFITM3 |
|---|---|
| Components |
|
-Supramolecule #1: A549 cell stably overexpressing IFITM3
| Supramolecule | Name: A549 cell stably overexpressing IFITM3 / type: cell / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
| Sectioning | Focused ion beam - Instrument: OTHER / Focused ion beam - Ion: OTHER / Focused ion beam - Voltage: 30 / Focused ion beam - Current: 0.3 / Focused ion beam - Duration: 600 / Focused ion beam - Temperature: 100 K / Focused ion beam - Initial thickness: 6000 / Focused ion beam - Final thickness: 150 Focused ion beam - Details: The value given for _em_focused_ion_beam.instrument is ThermoFisher Aquilos. This is not in a list of allowed values {'DB235', 'OTHER'} so OTHER is written into the XML file. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 3.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 0.004 µm / Nominal defocus min: 0.004 µm / Nominal magnification: 33000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | The tomogram was denoised using cryoCARE. |
|---|---|
| Final reconstruction | Software - Name: SerialEM / Number images used: 41 |
 Movie
Movie Controller
Controller