+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Influenza A/H7N9 polymerase apo-protein dimer complex | |||||||||
 マップデータ マップデータ | LocScale filtered map | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | Influenza / H7N9 / viral RNA-dependent RNA polymerase / VIRAL PROTEIN | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報cap snatching / viral transcription / symbiont-mediated suppression of host mRNA transcription via inhibition of RNA polymerase II activity / host cell mitochondrion / 7-methylguanosine mRNA capping / virion component / endonuclease activity / host cell cytoplasm / 加水分解酵素; エステル加水分解酵素 / symbiont-mediated suppression of host gene expression ...cap snatching / viral transcription / symbiont-mediated suppression of host mRNA transcription via inhibition of RNA polymerase II activity / host cell mitochondrion / 7-methylguanosine mRNA capping / virion component / endonuclease activity / host cell cytoplasm / 加水分解酵素; エステル加水分解酵素 / symbiont-mediated suppression of host gene expression / viral translational frameshifting / RNA-directed RNA polymerase / nucleotide binding / viral RNA genome replication / hydrolase activity / RNA-directed RNA polymerase activity / DNA-templated transcription / host cell nucleus / RNA binding / metal ion binding 類似検索 - 分子機能 | |||||||||
| 生物種 |  Influenza A virus (A/Zhejiang/DTID-ZJU01/2013(H7N9)) (A型インフルエンザウイルス) Influenza A virus (A/Zhejiang/DTID-ZJU01/2013(H7N9)) (A型インフルエンザウイルス) | |||||||||
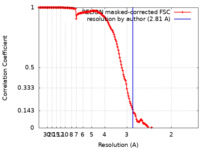
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.81 Å | |||||||||
 データ登録者 データ登録者 | Cusack S / Kouba T | |||||||||
| 資金援助 |  フランス, 1件 フランス, 1件
| |||||||||
 引用 引用 |  ジャーナル: Cell Rep / 年: 2023 ジャーナル: Cell Rep / 年: 2023タイトル: Direct observation of backtracking by influenza A and B polymerases upon consecutive incorporation of the nucleoside analog T1106. 著者: Tomas Kouba / Anna Dubankova / Petra Drncova / Elisa Donati / Pietro Vidossich / Valentina Speranzini / Alex Pflug / Johanna Huchting / Chris Meier / Marco De Vivo / Stephen Cusack /    要旨: The antiviral pseudo-base T705 and its de-fluoro analog T1106 mimic adenine or guanine and can be competitively incorporated into nascent RNA by viral RNA-dependent RNA polymerases. Although ...The antiviral pseudo-base T705 and its de-fluoro analog T1106 mimic adenine or guanine and can be competitively incorporated into nascent RNA by viral RNA-dependent RNA polymerases. Although dispersed, single pseudo-base incorporation is mutagenic, consecutive incorporation causes polymerase stalling and chain termination. Using a template encoding single and then consecutive T1106 incorporation four nucleotides later, we obtained a cryogenic electron microscopy structure of stalled influenza A/H7N9 polymerase. This shows that the entire product-template duplex backtracks by 5 nt, bringing the singly incorporated T1106 to the +1 position, where it forms an unexpected T1106:U wobble base pair. Similar structures show that influenza B polymerase also backtracks after consecutive T1106 incorporation, regardless of whether prior single incorporation has occurred. These results give insight into the unusual mechanism of chain termination by pyrazinecarboxamide base analogs. Consecutive incorporation destabilizes the proximal end of the product-template duplex, promoting irreversible backtracking to a more energetically favorable overall configuration. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_14858.map.gz emd_14858.map.gz | 159.4 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-14858-v30.xml emd-14858-v30.xml emd-14858.xml emd-14858.xml | 24.2 KB 24.2 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_14858_fsc.xml emd_14858_fsc.xml | 15.6 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_14858.png emd_14858.png | 75.7 KB | ||
| Filedesc metadata |  emd-14858.cif.gz emd-14858.cif.gz | 7.8 KB | ||
| その他 |  emd_14858_additional_1.map.gz emd_14858_additional_1.map.gz emd_14858_half_map_1.map.gz emd_14858_half_map_1.map.gz emd_14858_half_map_2.map.gz emd_14858_half_map_2.map.gz | 304.1 MB 257.3 MB 257.1 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14858 http://ftp.pdbj.org/pub/emdb/structures/EMD-14858 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14858 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14858 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  7zpmMC  7qtlC  7r0eC  7r1fC  7zplC  8bdrC  8be0C  8bekC  8bf5C M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_14858.map.gz / 形式: CCP4 / 大きさ: 325 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_14858.map.gz / 形式: CCP4 / 大きさ: 325 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | LocScale filtered map | ||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.8311 Å | ||||||||||||||||||||||||||||||||||||
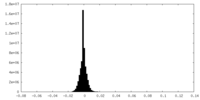
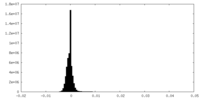
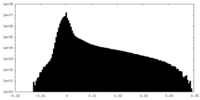
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-追加マップ: Relion post-processed map
| ファイル | emd_14858_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Relion post-processed map | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
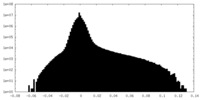
| 密度ヒストグラム |
-ハーフマップ: #2
| ファイル | emd_14858_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
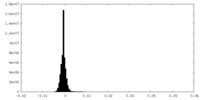
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_14858_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
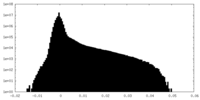
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : Influenza A/H7N9 polymerase apo-protein dimer complex
| 全体 | 名称: Influenza A/H7N9 polymerase apo-protein dimer complex |
|---|---|
| 要素 |
|
-超分子 #1: Influenza A/H7N9 polymerase apo-protein dimer complex
| 超分子 | 名称: Influenza A/H7N9 polymerase apo-protein dimer complex タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: all |
|---|---|
| 由来(天然) | 生物種:  Influenza A virus (A/Zhejiang/DTID-ZJU01/2013(H7N9)) (A型インフルエンザウイルス) Influenza A virus (A/Zhejiang/DTID-ZJU01/2013(H7N9)) (A型インフルエンザウイルス) |
| 分子量 | 理論値: 510 KDa |
-分子 #1: Polymerase acidic protein
| 分子 | 名称: Polymerase acidic protein / タイプ: protein_or_peptide / ID: 1 / コピー数: 2 / 光学異性体: LEVO EC番号: 加水分解酵素; エステル加水分解酵素 |
|---|---|
| 由来(天然) | 生物種:  Influenza A virus (A/Zhejiang/DTID-ZJU01/2013(H7N9)) (A型インフルエンザウイルス) Influenza A virus (A/Zhejiang/DTID-ZJU01/2013(H7N9)) (A型インフルエンザウイルス)株: A/Zhejiang/DTID-ZJU01/2013(H7N9) |
| 分子量 | 理論値: 82.930602 KDa |
| 組換発現 | 生物種:  Trichoplusia ni (イラクサキンウワバ) Trichoplusia ni (イラクサキンウワバ) |
| 配列 | 文字列: GMEDFVRQCF NPMIVELAEK AMKEYGEDPK IETNKFASIC THLEVCFMYS DFHFIDERGE STIIESGDPN VLLKHRFEII EGRDRTMAW TVVNSICNTT GVEKPKFLPD LYDYKENRFI EIGVTRREVH IYYLEKANKI KSEKTHIHIF SFTGEEMATK A DYTLDEES ...文字列: GMEDFVRQCF NPMIVELAEK AMKEYGEDPK IETNKFASIC THLEVCFMYS DFHFIDERGE STIIESGDPN VLLKHRFEII EGRDRTMAW TVVNSICNTT GVEKPKFLPD LYDYKENRFI EIGVTRREVH IYYLEKANKI KSEKTHIHIF SFTGEEMATK A DYTLDEES RARIKTRLFT IRQEMASRGL WDSFRQSERG EETIEERFEI TGTMRRLADQ SLPPNFSSLE NFRAYVDGFE PN GCIEGKL SQMSKEVNAR IEPFLRTTPR PLRLPDGPPC SQRSKFLLMD ALKLSIEDPS HEGEGIPLYD AIKCMKTFFG WKE PNIIKP HEKGINPNYL LTWKQVLAEL QDIENEEKIP RTKNMKKTSQ LKWALGENMA PEKVDFEDCK DVNDLKQYDS DEPE PRSLA CWIQSEFNKA CELTDSSWVE LDEIGEDVAP IEHIASMRRN YFTAEVSHCR ATEYIMKGVY INTALLNASC AAMDD FQLI PMISKCRTKE GRRKTNLYGF IIKGRSHLRN DTDVVNFVSM EFSLTDPRLE PHKWEKYCVL EIGDMLLRTA VGQVSR PMF LYVRTNGTSK IKMKWGMEMR RCLLQSLQQI ESMIEAESSV KEKDLTKEFF ENKSETWPIG ESPKGVEEGS IGKVCRT LL AKSVFNSLYA SPQLEGFSAE SRKLLLIVQA LRDNLEPGTF DLEGLYEAIE ECLINDPWVL LNASWFNSFL THALR UniProtKB: Polymerase acidic protein |
-分子 #2: RNA-directed RNA polymerase catalytic subunit
| 分子 | 名称: RNA-directed RNA polymerase catalytic subunit / タイプ: protein_or_peptide / ID: 2 / コピー数: 2 / 光学異性体: LEVO / EC番号: RNA-directed RNA polymerase |
|---|---|
| 由来(天然) | 生物種:  Influenza A virus (A/Zhejiang/DTID-ZJU01/2013(H7N9)) (A型インフルエンザウイルス) Influenza A virus (A/Zhejiang/DTID-ZJU01/2013(H7N9)) (A型インフルエンザウイルス)株: A/Zhejiang/DTID-ZJU01/2013(H7N9) |
| 分子量 | 理論値: 86.496156 KDa |
| 組換発現 | 生物種:  Trichoplusia ni (イラクサキンウワバ) Trichoplusia ni (イラクサキンウワバ) |
| 配列 | 文字列: MDVNPTLLFL KVPVQNAIST TFPYTGDPPY SHGTGTGYTM DTVNRTHKYS EKGKWTTNTE TGAPQLNPID GPLPEDNEPS GYAQTDCVL EAMAFLEESH PGIFENSCLE TMEIVQQTRV DKLTQGRQTY DWTLNRNQPA ATALANTIEV FRSNGLTANE S GRLIDFLK ...文字列: MDVNPTLLFL KVPVQNAIST TFPYTGDPPY SHGTGTGYTM DTVNRTHKYS EKGKWTTNTE TGAPQLNPID GPLPEDNEPS GYAQTDCVL EAMAFLEESH PGIFENSCLE TMEIVQQTRV DKLTQGRQTY DWTLNRNQPA ATALANTIEV FRSNGLTANE S GRLIDFLK DVMDSMDKEE MEITTHFQRK RRVRDNMTKK MVTQRTIGKK KQRLNKRSYL IRALTLNTMT KDAERGKLKR RA IATPGMQ IRGFVYFVEA LARSICEKLE QSGLPVGGNE KKAKLANVVR KMMTNSQDTE LSFTITGDNT KWNENQNPRM FLA MITYIT RNQPEWFRNV LSIAPIMFSN KMARLGKGYM FESKSMKLRT QVPAEMLANI DLKYFNKSTR EKIEKIRPLL IDGT ASLSP GMMMGMFNML STVLGVSILN LGQKKYTKTT YWWDGLQSSD DFALIVNAPN HEGIQAGVDR FYRTCKLVGI NMSKK KSYI NRTGTFEFTS FFYRYGFVAN FSMELPSFGV SGINESADMS VGVTVIKNNM INNDLGPATA QMALQLFIKD YRYTYR CHR GDTQIQTRRA FELKKLWEQT RSKAGLLVSD GGPNLYNIRN LHIPEVCLKW ELMDEDYQGR LCNPMNPFVS HKEIDSV NN AVVMPAHGPA KSMEYDAVAT THSWIPKRNR SILNTSQRGI LEDEQMYQKC CNLFEKFFPS SSYRRPVGIS SMVEAMVS R ARIDARIDFE SGRIKKEEFA EIMKICSTIE ELRRQK UniProtKB: RNA-directed RNA polymerase catalytic subunit |
-分子 #3: Polymerase basic protein 2
| 分子 | 名称: Polymerase basic protein 2 / タイプ: protein_or_peptide / ID: 3 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Influenza A virus (A/Zhejiang/DTID-ZJU01/2013(H7N9)) (A型インフルエンザウイルス) Influenza A virus (A/Zhejiang/DTID-ZJU01/2013(H7N9)) (A型インフルエンザウイルス)株: A/Zhejiang/DTID-ZJU01/2013(H7N9) |
| 分子量 | 理論値: 86.00732 KDa |
| 組換発現 | 生物種:  Trichoplusia ni (イラクサキンウワバ) Trichoplusia ni (イラクサキンウワバ) |
| 配列 | 文字列: MERIKELRDL MSQSRTREIL TKTTVDHMAI IKKYTSGRQE KNPALRMKWM MAMKYPITAD KRIMEMIPER NEQGQTLWSK TNDAGSDRV MVSPLAVTWW NRNGPTTSTV HYPKVYKTYF EKVERLKHGT FGPVHFRNQV KIRRRVDINP GHADLSAKEA Q DVIMEVVF ...文字列: MERIKELRDL MSQSRTREIL TKTTVDHMAI IKKYTSGRQE KNPALRMKWM MAMKYPITAD KRIMEMIPER NEQGQTLWSK TNDAGSDRV MVSPLAVTWW NRNGPTTSTV HYPKVYKTYF EKVERLKHGT FGPVHFRNQV KIRRRVDINP GHADLSAKEA Q DVIMEVVF PNEVGARILT SESQLTITKE KKKELQDCKI APLMVAYMLE RELVRKTRFL PVAGGTSSVY IEVLHLTQGT CW EQMYTPG GEVRNDDVDQ SLIIAARNIV RRATVSADPL ASLLEMCHST QIGGVRMVDI LRQNPTEEQA VDICKAAMGL RIS SSFSFG GFTFKRTSGS SVKREEEVLT GNLQTLKIRV HEGYEEFTMV GRRATAILRK ATRRLIQLIV SGKDEQSIAE AIIV AMVFS QEDCMIKAVR GDLNFVNRAN QRLNPMHQLL RHFQKDAKVL FQNWGIEPID NVMGMIGILP DMTPSTEMSL RGVRV SKMG VDEYSSTERV VVSIDRFLRV RDQRGNVLLS PEEVSETQGT EKLTITYSSS MMWEINGPES VLVNTYQWII RNWENV KIQ WSQDPTMLYN KMEFEPFQSL VPKAARGQYS GFVRVLFQQM RDVLGTFDTV QIIKLLPFAA APPEQSRMQF SSLTVNV RG SGMRIVVRGN SPVFNYNKAT KRLTVLGKDA GALMEDPDEG TAGVESAVLR GFLILGKENK RYGPALSINE LSNLAKGE K ANVLIGQGDV VLVMKRKRDS SILTDSQTAT KRIRMAIN UniProtKB: Polymerase basic protein 2 |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | TFS KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 QUANTUM (4k x 4k) 平均電子線量: 48.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 3.0 µm / 最小 デフォーカス(公称値): 0.5 µm |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)