[English] 日本語
 Yorodumi
Yorodumi- EMDB-14023: In vitro assembled tau filaments into Quadruple Helical Filaments... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-14023 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | In vitro assembled tau filaments into Quadruple Helical Filaments type 1 (2c) | |||||||||
 Map data Map data | Sharpened map of in vitro assembled tau filaments into Quadruple Helical Filaments type 1 (2c) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Neurodegeneration / Amyloid / Tau / Alzheimer's Disease / PROTEIN FIBRIL | |||||||||
| Function / homology |  Function and homology information Function and homology informationplus-end-directed organelle transport along microtubule / histone-dependent DNA binding / negative regulation of protein localization to mitochondrion / neurofibrillary tangle / microtubule lateral binding / axonal transport / tubulin complex / positive regulation of protein localization to synapse / phosphatidylinositol bisphosphate binding / generation of neurons ...plus-end-directed organelle transport along microtubule / histone-dependent DNA binding / negative regulation of protein localization to mitochondrion / neurofibrillary tangle / microtubule lateral binding / axonal transport / tubulin complex / positive regulation of protein localization to synapse / phosphatidylinositol bisphosphate binding / generation of neurons / rRNA metabolic process / axonal transport of mitochondrion / regulation of mitochondrial fission / axon development / regulation of microtubule-based movement / regulation of chromosome organization / intracellular distribution of mitochondria / central nervous system neuron development / minor groove of adenine-thymine-rich DNA binding / lipoprotein particle binding / microtubule polymerization / negative regulation of mitochondrial membrane potential / regulation of microtubule polymerization / dynactin binding / apolipoprotein binding / protein polymerization / main axon / axolemma / Caspase-mediated cleavage of cytoskeletal proteins / regulation of microtubule polymerization or depolymerization / negative regulation of mitochondrial fission / glial cell projection / neurofibrillary tangle assembly / positive regulation of axon extension / positive regulation of microtubule polymerization / regulation of cellular response to heat / Activation of AMPK downstream of NMDARs / positive regulation of protein localization / positive regulation of superoxide anion generation / cellular response to brain-derived neurotrophic factor stimulus / regulation of long-term synaptic depression / supramolecular fiber organization / cytoplasmic microtubule organization / regulation of calcium-mediated signaling / somatodendritic compartment / axon cytoplasm / synapse assembly / astrocyte activation / phosphatidylinositol binding / nuclear periphery / enzyme inhibitor activity / protein phosphatase 2A binding / stress granule assembly / regulation of microtubule cytoskeleton organization / regulation of autophagy / cellular response to reactive oxygen species / microglial cell activation / cellular response to nerve growth factor stimulus / Hsp90 protein binding / protein homooligomerization / SH3 domain binding / PKR-mediated signaling / regulation of synaptic plasticity / synapse organization / response to lead ion / microtubule cytoskeleton organization / memory / cytoplasmic ribonucleoprotein granule / neuron projection development / cell-cell signaling / single-stranded DNA binding / cellular response to heat / growth cone / protein-folding chaperone binding / microtubule cytoskeleton / actin binding / cell body / double-stranded DNA binding / microtubule binding / sequence-specific DNA binding / amyloid fibril formation / dendritic spine / microtubule / protein-macromolecule adaptor activity / learning or memory / neuron projection / membrane raft / negative regulation of gene expression / axon / neuronal cell body / DNA damage response / dendrite / protein kinase binding / enzyme binding / mitochondrion / DNA binding / RNA binding / extracellular region / identical protein binding / nucleus Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
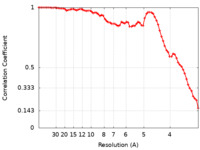
| Method | helical reconstruction / cryo EM / Resolution: 3.29 Å | |||||||||
 Authors Authors | Lovestam S / Scheres SHW | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2022 Journal: Elife / Year: 2022Title: Assembly of recombinant tau into filaments identical to those of Alzheimer's disease and chronic traumatic encephalopathy. Authors: Sofia Lövestam / Fujiet Adrian Koh / Bart van Knippenberg / Abhay Kotecha / Alexey G Murzin / Michel Goedert / Sjors H W Scheres /   Abstract: Abundant filamentous inclusions of tau are characteristic of more than 20 neurodegenerative diseases that are collectively termed tauopathies. Electron cryo-microscopy (cryo-EM) structures of tau ...Abundant filamentous inclusions of tau are characteristic of more than 20 neurodegenerative diseases that are collectively termed tauopathies. Electron cryo-microscopy (cryo-EM) structures of tau amyloid filaments from human brain revealed that distinct tau folds characterise many different diseases. A lack of laboratory-based model systems to generate these structures has hampered efforts to uncover the molecular mechanisms that underlie tauopathies. Here, we report in vitro assembly conditions with recombinant tau that replicate the structures of filaments from both Alzheimer's disease (AD) and chronic traumatic encephalopathy (CTE), as determined by cryo-EM. Our results suggest that post-translational modifications of tau modulate filament assembly, and that previously observed additional densities in AD and CTE filaments may arise from the presence of inorganic salts, like phosphates and sodium chloride. In vitro assembly of tau into disease-relevant filaments will facilitate studies to determine their roles in different diseases, as well as the development of compounds that specifically bind to these structures or prevent their formation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14023.map.gz emd_14023.map.gz | 9.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14023-v30.xml emd-14023-v30.xml emd-14023.xml emd-14023.xml | 15.9 KB 15.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14023_fsc.xml emd_14023_fsc.xml | 6.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_14023.png emd_14023.png | 47 KB | ||
| Filedesc metadata |  emd-14023.cif.gz emd-14023.cif.gz | 5.4 KB | ||
| Others |  emd_14023_additional_1.map.gz emd_14023_additional_1.map.gz emd_14023_half_map_1.map.gz emd_14023_half_map_1.map.gz emd_14023_half_map_2.map.gz emd_14023_half_map_2.map.gz | 20.4 MB 20.6 MB 20.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14023 http://ftp.pdbj.org/pub/emdb/structures/EMD-14023 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14023 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14023 | HTTPS FTP |
-Related structure data
| Related structure data |  7qjvMC  7qjwC  7qjxC  7qjyC  7qjzC  7qk1C  7qk2C  7qk3C  7qk5C  7qk6C  7qkfC  7qkgC  7qkhC  7qkiC  7qkjC  7qkkC  7qklC  7qkmC  7qkuC  7qkvC  7qkwC  7qkxC  7qkyC  7qkzC  7ql0C  7ql1C  7ql2C  7ql3C  7ql4C  7r4tC  7r5hC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14023.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14023.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map of in vitro assembled tau filaments into Quadruple Helical Filaments type 1 (2c) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.648 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
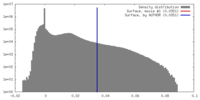
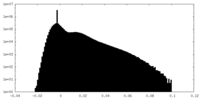
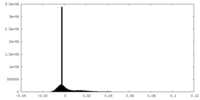
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Map of in vitro assembled tau filaments into...
| File | emd_14023_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map of in vitro assembled tau filaments into Quadruple Helical Filaments type 1 (2c) | ||||||||||||
| Projections & Slices |
| ||||||||||||
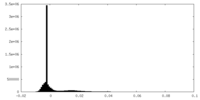
| Density Histograms |
-Half map: Half map 1 of in vitro assembled tau...
| File | emd_14023_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 of in vitro assembled tau filaments into Quadruple Helical Filaments type 1 (2c) | ||||||||||||
| Projections & Slices |
| ||||||||||||
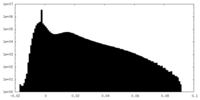
| Density Histograms |
-Half map: Half map 2 of in vitro assembled tau...
| File | emd_14023_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 of in vitro assembled tau filaments into Quadruple Helical Filaments type 1 (2c) | ||||||||||||
| Projections & Slices |
| ||||||||||||
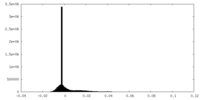
| Density Histograms |
- Sample components
Sample components
-Entire : Amyloid filament
| Entire | Name: Amyloid filament |
|---|---|
| Components |
|
-Supramolecule #1: Amyloid filament
| Supramolecule | Name: Amyloid filament / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Microtubule-associated protein tau
| Macromolecule | Name: Microtubule-associated protein tau / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 45.919871 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAEPRQEFEV MEDHAGTYGL GDRKDQGGYT MHQDQEGDTD AGLKESPLQT PTEDGSEEPG SETSDAKSTP TAEDVTAPLV DEGAPGKQA AAQPHTEIPE GTTAEEAGIG DTPSLEDEAA GHVTQARMVS KSKDGTGSDD KKAKGADGKT KIATPRGAAP P GQKGQANA ...String: MAEPRQEFEV MEDHAGTYGL GDRKDQGGYT MHQDQEGDTD AGLKESPLQT PTEDGSEEPG SETSDAKSTP TAEDVTAPLV DEGAPGKQA AAQPHTEIPE GTTAEEAGIG DTPSLEDEAA GHVTQARMVS KSKDGTGSDD KKAKGADGKT KIATPRGAAP P GQKGQANA TRIPAKTPPA PKTPPSSGEP PKSGDRSGYS SPGSPGTPGS RSRTPSLPTP PTREPKKVAV VRTPPKSPSS AK SRLQTAP VPMPDLKNVK SKIGSTENLK HQPGGGKVQI INKKLDLSNV QSKCGSKDNI KHVPGGGSVQ IVYKPVDLSK VTS KCGSLG NIHHKPGGGQ VEVKSEKLDF KDRVQSKIGS LDNITHVPGG GNKKIETHKL TFRENAKAKT DHGAEIVYKS PVVS GDTSP RHLSNVSSTG SIDMVDSPQL ATLADEVSAS LAKQGL UniProtKB: Microtubule-associated protein tau |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)