[English] 日本語
 Yorodumi
Yorodumi- EMDB-13899: Structure of the MUCIN-2 Cterminal domains partially deglycosylated. -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the MUCIN-2 Cterminal domains partially deglycosylated. | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Mucus / extracellular / net assemble / STRUCTURAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationinner mucus layer / outer mucus layer / host-mediated modulation of intestinal microbiota composition / Defective GALNT3 causes HFTC / Defective C1GALT1C1 causes TNPS / Defective GALNT12 causes CRCS1 / Termination of O-glycan biosynthesis / O-linked glycosylation of mucins / maintenance of gastrointestinal epithelium / mucus secretion ...inner mucus layer / outer mucus layer / host-mediated modulation of intestinal microbiota composition / Defective GALNT3 causes HFTC / Defective C1GALT1C1 causes TNPS / Defective GALNT12 causes CRCS1 / Termination of O-glycan biosynthesis / O-linked glycosylation of mucins / maintenance of gastrointestinal epithelium / mucus secretion / detoxification of copper ion / cupric ion binding / Dectin-2 family / cuprous ion binding / Golgi lumen / extracellular matrix / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.25 Å | |||||||||
 Authors Authors | Gallego P / Hansson GC | |||||||||
| Funding support |  Sweden, 1 items Sweden, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: The intestinal MUC2 mucin C-terminus is stabilized by an extra disulfide bond in comparison to von Willebrand factor and other gel-forming mucins. Authors: Pablo Gallego / Maria-Jose Garcia-Bonete / Sergio Trillo-Muyo / Christian V Recktenwald / Malin E V Johansson / Gunnar C Hansson /  Abstract: The MUC2 mucin polymer is the main building unit of the intestinal mucus layers separating intestinal microbiota from the host epithelium. The MUC2 mucin is a large glycoprotein with a C-terminal ...The MUC2 mucin polymer is the main building unit of the intestinal mucus layers separating intestinal microbiota from the host epithelium. The MUC2 mucin is a large glycoprotein with a C-terminal domain similar to the MUC5AC and MUC5B mucins and the von Willebrand factor (VWF). A structural model of the C-terminal part of MUC2, MUC2-C, was generated by combining Cryo-electron microscopy, AlphaFold prediction, information of its glycosylation, and small angle X-ray scattering information. The globular VWD4 assembly in the N-terminal of MUC2-C is followed by 3.5 linear VWC domains that form an extended flexible structure before the C-terminal cystine-knot. All gel-forming mucins and VWF form tail-tail disulfide-bonded dimers in their C-terminal cystine-knot domain, but interestingly the MUC2 mucin has an extra stabilizing disulfide bond on the N-terminal side of the VWD4 domain, likely essential for a stable intestinal mucus barrier. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13899.map.gz emd_13899.map.gz | 153 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13899-v30.xml emd-13899-v30.xml emd-13899.xml emd-13899.xml | 12.1 KB 12.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_13899.png emd_13899.png | 27.1 KB | ||
| Filedesc metadata |  emd-13899.cif.gz emd-13899.cif.gz | 6.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13899 http://ftp.pdbj.org/pub/emdb/structures/EMD-13899 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13899 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13899 | HTTPS FTP |
-Related structure data
| Related structure data |  7qcuMC  7qclC  7qcnC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13899.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13899.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.82 Å | ||||||||||||||||||||||||||||||||||||
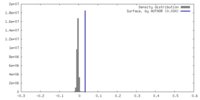
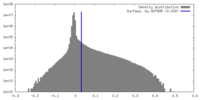
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Muc2 dimer, Mucus assemble
| Entire | Name: Muc2 dimer, Mucus assemble |
|---|---|
| Components |
|
-Supramolecule #1: Muc2 dimer, Mucus assemble
| Supramolecule | Name: Muc2 dimer, Mucus assemble / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Mucin-2
| Macromolecule | Name: Mucin-2 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 88.659469 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AAQPARRAVR SSRRHHHHHH GSGLEVLFQG PTPGTKPPEC PDFDPPRQEN ETWWLCDCFM ATCKYNNTVE IVKVECEPPP MPTCSNGLQ PVRVEDPDGC CWHWECDCYC TGWGDPHYVT FDGLYYSYQG NCTYVLVEEI SPSVDNFGVY IDNYHCDPND K VSCPRTLI ...String: AAQPARRAVR SSRRHHHHHH GSGLEVLFQG PTPGTKPPEC PDFDPPRQEN ETWWLCDCFM ATCKYNNTVE IVKVECEPPP MPTCSNGLQ PVRVEDPDGC CWHWECDCYC TGWGDPHYVT FDGLYYSYQG NCTYVLVEEI SPSVDNFGVY IDNYHCDPND K VSCPRTLI VRHETQEVLI KTVHMMPMQV QVQVNRQAVA LPYKKYGLEV YQSGINYVVD IPELGVLVSY NGLSFSVRLP YH RFGNNTK GQCGTCTNTT SDDCILPSGE IVSNCEAAAD QWLVNDPSKP HCPHSSSTTK RPAVTVPGGG KTTPHKDCTP SPL CQLIKD SLFAQCHALV PPQHYYDACV FDSCFMPGSS LECASLQAYA ALCAQQNICL DWRNHTHGAC LVECPSHREY QACG PAEEP TCKSSSSQQN NTVLVEGCFC PEGTMNYAPG FDVCVKTCGC VGPDNVPREF GEHFEFDCKN CVCLEGGSGI ICQPK RCSQ KPVTHCVEDG TYLATEVNPA DTCCNITVCK CNTSLCKEKP SVCPLGFEVK SKMVPGRCCP FYWCESKGVC VHGNAE YQP GSPVYSSKCQ DCVCTDKVDN NTLLNVIACT HVPCNTSCSP GFELMEAPGE CCKKCEQTHC IIKRPDNQHV ILKPGDF KS DPKNNCTFFS CVKIHNQLIS SVSNITCPNF DASICIPGSI TFMPNGCCKT CTPRNETRVP CSTVPVTTEV SYAGCTKT V LMNHCSGSCG TFVMYSAKAQ ALDHSCSCCK EEKTSQREVV LSCPNGGSLT HTYTHIESCQ CQDTVCGLPT GTSRRARRS PRHLGSG UniProtKB: Mucin-2 |
-Macromolecule #3: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 3 / Number of copies: 2 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 8 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 79.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.25 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. 3) / Number images used: 387288 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 3) |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-7qcu: |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)