+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the plectasin fibril (double strands) | |||||||||
 Map data Map data | Plectasin double strands | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | fibril / helical / ANTIMICROBIAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationpotassium channel regulator activity / toxin activity / defense response to bacterium / host cell plasma membrane / extracellular region Similarity search - Function | |||||||||
| Biological species |  Pseudoplectania nigrella (fungus) Pseudoplectania nigrella (fungus) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 2.0 Å | |||||||||
 Authors Authors | Effantin G | |||||||||
| Funding support | European Union, 1 items
| |||||||||
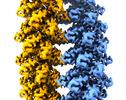
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: pH- and concentration-dependent supramolecular assembly of a fungal defensin plectasin variant into helical non-amyloid fibrils. Authors: Christin Pohl / Gregory Effantin / Eaazhisai Kandiah / Sebastian Meier / Guanghong Zeng / Werner Streicher / Dorotea Raventos Segura / Per H Mygind / Dorthe Sandvang / Line Anker Nielsen / ...Authors: Christin Pohl / Gregory Effantin / Eaazhisai Kandiah / Sebastian Meier / Guanghong Zeng / Werner Streicher / Dorotea Raventos Segura / Per H Mygind / Dorthe Sandvang / Line Anker Nielsen / Günther H J Peters / Guy Schoehn / Christoph Mueller-Dieckmann / Allan Noergaard / Pernille Harris /     Abstract: Self-assembly and fibril formation play important roles in protein behaviour. Amyloid fibril formation is well-studied due to its role in neurodegenerative diseases and characterized by refolding of ...Self-assembly and fibril formation play important roles in protein behaviour. Amyloid fibril formation is well-studied due to its role in neurodegenerative diseases and characterized by refolding of the protein into predominantly β-sheet form. However, much less is known about the assembly of proteins into other types of supramolecular structures. Using cryo-electron microscopy at a resolution of 1.97 Å, we show that a triple-mutant of the anti-microbial peptide plectasin, PPI42, assembles into helical non-amyloid fibrils. The in vitro anti-microbial activity was determined and shown to be enhanced compared to the wildtype. Plectasin contains a cysteine-stabilised α-helix-β-sheet structure, which remains intact upon fibril formation. Two protofilaments form a right-handed protein fibril. The fibril formation is reversible and follows sigmoidal kinetics with a pH- and concentration dependent equilibrium between soluble monomer and protein fibril. This high-resolution structure reveals that α/β proteins can natively assemble into fibrils. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12775.map.gz emd_12775.map.gz | 78.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12775-v30.xml emd-12775-v30.xml emd-12775.xml emd-12775.xml | 12.4 KB 12.4 KB | Display Display |  EMDB header EMDB header |
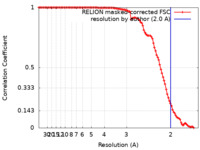
| FSC (resolution estimation) |  emd_12775_fsc.xml emd_12775_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_12775.png emd_12775.png | 175.1 KB | ||
| Filedesc metadata |  emd-12775.cif.gz emd-12775.cif.gz | 5.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12775 http://ftp.pdbj.org/pub/emdb/structures/EMD-12775 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12775 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12775 | HTTPS FTP |
-Related structure data
| Related structure data |  7oaeMC  7o76C  7oagC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12775.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12775.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Plectasin double strands | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.827 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : supermolecular assembly of plectasin into two protein fibrils
| Entire | Name: supermolecular assembly of plectasin into two protein fibrils |
|---|---|
| Components |
|
-Supramolecule #1: supermolecular assembly of plectasin into two protein fibrils
| Supramolecule | Name: supermolecular assembly of plectasin into two protein fibrils type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Pseudoplectania nigrella (fungus) Pseudoplectania nigrella (fungus) |
-Macromolecule #1: Fungal defensin plectasin
| Macromolecule | Name: Fungal defensin plectasin / type: protein_or_peptide / ID: 1 / Number of copies: 42 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudoplectania nigrella (fungus) Pseudoplectania nigrella (fungus) |
| Molecular weight | Theoretical: 4.402092 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GFGCNGPWSE DDMKCHNHCK SIKGYKGGYC AKGGFLCKCY UniProtKB: Fungal defensin plectasin |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Buffer | pH: 5.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)