+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12765 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Encequidar-bound human P-glycoprotein in complex with UIC2-Fab | |||||||||
 Map data Map data | encequidar-bound human P-glycoprotein in complex with UIC2-Fab | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of anion channel activity / carboxylic acid transmembrane transport / carboxylic acid transmembrane transporter activity / ceramide translocation / terpenoid transport / ceramide floppase activity / regulation of response to osmotic stress / floppase activity / Abacavir transmembrane transport / ABC-type oligopeptide transporter activity ...positive regulation of anion channel activity / carboxylic acid transmembrane transport / carboxylic acid transmembrane transporter activity / ceramide translocation / terpenoid transport / ceramide floppase activity / regulation of response to osmotic stress / floppase activity / Abacavir transmembrane transport / ABC-type oligopeptide transporter activity / external side of apical plasma membrane / Atorvastatin ADME / phosphatidylethanolamine flippase activity / xenobiotic transport across blood-brain barrier / phosphatidylcholine floppase activity / transepithelial transport / xenobiotic detoxification by transmembrane export across the plasma membrane / export across plasma membrane / ABC-type xenobiotic transporter / P-type phospholipid transporter / ABC-type xenobiotic transporter activity / phospholipid translocation / Prednisone ADME / efflux transmembrane transporter activity / xenobiotic transmembrane transporter activity / transmembrane transporter activity / transport across blood-brain barrier / ATPase-coupled transmembrane transporter activity / xenobiotic metabolic process / regulation of chloride transport / stem cell proliferation / ABC-family proteins mediated transport / transmembrane transport / G2/M transition of mitotic cell cycle / response to xenobiotic stimulus / apical plasma membrane / ubiquitin protein ligase binding / cell surface / ATP hydrolysis activity / extracellular exosome / ATP binding / membrane / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
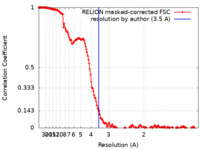
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Nosol K / Locher KP | |||||||||
| Funding support |  Switzerland, 1 items Switzerland, 1 items
| |||||||||
 Citation Citation |  Journal: J Med Chem / Year: 2022 Journal: J Med Chem / Year: 2022Title: Discovery and Characterization of Potent Dual P-Glycoprotein and CYP3A4 Inhibitors: Design, Synthesis, Cryo-EM Analysis, and Biological Evaluations. Authors: Sameer Urgaonkar / Kamil Nosol / Ahmed M Said / Nader N Nasief / Yahao Bu / Kaspar P Locher / Johnson Y N Lau / Michael P Smolinski /   Abstract: Targeted concurrent inhibition of intestinal drug efflux transporter P-glycoprotein (P-gp) and drug metabolizing enzyme cytochrome P450 3A4 (CYP3A4) is a promising approach to improve oral ...Targeted concurrent inhibition of intestinal drug efflux transporter P-glycoprotein (P-gp) and drug metabolizing enzyme cytochrome P450 3A4 (CYP3A4) is a promising approach to improve oral bioavailability of their common substrates such as docetaxel, while avoiding side effects arising from their pan inhibitions. Herein, we report the discovery and characterization of potent small molecule inhibitors of P-gp and CYP3A4 with encequidar (minimally absorbed P-gp inhibitor) as a starting point for optimization. To aid in the design of these dual inhibitors, we solved the high-resolution cryo-EM structure of encequidar bound to human P-gp. The structure guided us to prudently decorate the encequidar scaffold with CYP3A4 pharmacophores, leading to the identification of several analogues with dual potency against P-gp and CYP3A4. , dual P-gp and CYP3A4 inhibitor improved the oral absorption of docetaxel by 3-fold as compared to vehicle, while itself remained poorly absorbed. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12765.map.gz emd_12765.map.gz | 320.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12765-v30.xml emd-12765-v30.xml emd-12765.xml emd-12765.xml | 14.9 KB 14.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12765_fsc.xml emd_12765_fsc.xml | 16 KB | Display |  FSC data file FSC data file |
| Images |  emd_12765.png emd_12765.png | 105.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12765 http://ftp.pdbj.org/pub/emdb/structures/EMD-12765 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12765 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12765 | HTTPS FTP |
-Validation report
| Summary document |  emd_12765_validation.pdf.gz emd_12765_validation.pdf.gz | 396.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_12765_full_validation.pdf.gz emd_12765_full_validation.pdf.gz | 395.8 KB | Display | |
| Data in XML |  emd_12765_validation.xml.gz emd_12765_validation.xml.gz | 14.7 KB | Display | |
| Data in CIF |  emd_12765_validation.cif.gz emd_12765_validation.cif.gz | 20.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12765 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12765 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12765 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12765 | HTTPS FTP |
-Related structure data
| Related structure data |  7o9wMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12765.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12765.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | encequidar-bound human P-glycoprotein in complex with UIC2-Fab | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.66 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Nanodisc reconstituted human P-glycoprotein in complex with UIC2-...
| Entire | Name: Nanodisc reconstituted human P-glycoprotein in complex with UIC2-Fab and encequidar |
|---|---|
| Components |
|
-Supramolecule #1: Nanodisc reconstituted human P-glycoprotein in complex with UIC2-...
| Supramolecule | Name: Nanodisc reconstituted human P-glycoprotein in complex with UIC2-Fab and encequidar type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Molecular weight | Experimental: 240 KDa |
-Supramolecule #2: Multidrug resistance protein 1
| Supramolecule | Name: Multidrug resistance protein 1 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Fab
| Supramolecule | Name: Fab / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Multidrug resistance protein 1
| Macromolecule | Name: Multidrug resistance protein 1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: ABC-type xenobiotic transporter |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 141.628781 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDLEGDRNGG AKKKNFFKLN NKSEKDKKEK KPTVSVFSMF RYSNWLDKLY MVVGTLAAII HGAGLPLMML VFGEMTDIFA NAGNLEDLM SNITNRSDIN DTGFFMNLEE DMTRYAYYYS GIGAGVLVAA YIQVSFWCLA AGRQIHKIRK QFFHAIMRQE I GWFDVHDV ...String: MDLEGDRNGG AKKKNFFKLN NKSEKDKKEK KPTVSVFSMF RYSNWLDKLY MVVGTLAAII HGAGLPLMML VFGEMTDIFA NAGNLEDLM SNITNRSDIN DTGFFMNLEE DMTRYAYYYS GIGAGVLVAA YIQVSFWCLA AGRQIHKIRK QFFHAIMRQE I GWFDVHDV GELNTRLTDD VSKINEGIGD KIGMFFQSMA TFFTGFIVGF TRGWKLTLVI LAISPVLGLS AAVWAKILSS FT DKELLAY AKAGAVAEEV LAAIRTVIAF GGQKKELERY NKNLEEAKRI GIKKAITANI SIGAAFLLIY ASYALAFWYG TTL VLSGEY SIGQVLTVFF SVLIGAFSVG QASPSIEAFA NARGAAYEIF KIIDNKPSID SYSKSGHKPD NIKGNLEFRN VHFS YPSRK EVKILKGLNL KVQSGQTVAL VGNSGCGKST TVQLMQRLYD PTEGMVSVDG QDIRTINVRF LREIIGVVSQ EPVLF ATTI AENIRYGREN VTMDEIEKAV KEANAYDFIM KLPHKFDTLV GERGAQLSGG QKQRIAIARA LVRNPKILLL DEATSA LDT ESEAVVQVAL DKARKGRTTI VIAHRLSTVR NADVIAGFDD GVIVEKGNHD ELMKEKGIYF KLVTMQTAGN EVELENA AD ESKSEIDALE MSSNDSRSSL IRKRSTRRSV RGSQAQDRKL STKEALDESI PPVSFWRIMK LNLTEWPYFV VGVFCAII N GGLQPAFAII FSKIIGVFTR IDDPETKRQN SNLFSLLFLA LGIISFITFF LQGFTFGKAG EILTKRLRYM VFRSMLRQD VSWFDDPKNT TGALTTRLAN DAAQVKGAIG SRLAVITQNI ANLGTGIIIS FIYGWQLTLL LLAIVPIIAI AGVVEMKMLS GQALKDKKE LEGAGKIATE AIENFRTVVS LTQEQKFEHM YAQSLQVPYR NSLRKAHIFG ITFSFTQAMM YFSYAGCFRF G AYLVAHKL MSFEDVLLVF SAVVFGAMAV GQVSSFAPDY AKAKISAAHI IMIIEKTPLI DSYSTEGLMP NTLEGNVTFG EV VFNYPTR PDIPVLQGLS LEVKKGQTLA LVGSSGCGKS TVVQLLERFY DPLAGKVLLD GKEIKRLNVQ WLRAHLGIVS QEP ILFDCS IAENIAYGDN SRVVSQEEIV RAAKEANIHA FIESLPNKYS TKVGDKGTQL SGGQKQRIAI ARALVRQPHI LLLD EATSA LDTESEKVVQ EALDKAREGR TCIVIAHRLS TIQNADLIVV FQNGRVKEHG THQQLLAQKG IYFSMVSVQA GTKRQ |
-Macromolecule #2: UIC2 Fab-fragment light chain
| Macromolecule | Name: UIC2 Fab-fragment light chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 24.321039 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVVMTQSPLS LPVSLGDQAS ISCRSSQSLL HSNGNTYLHW YLQKPGQSPK LLIYKVSNRF SGVPDRFSGS GSGTDFTLKI SRVEAEDLG VYFCSQSTHI PPWTFGGGTK LDIKRADAAP TVSIFPPSSE QLTSGGLSVV CFLNNFYPKD INVKWKIDGS E RQNGVLNS ...String: QVVMTQSPLS LPVSLGDQAS ISCRSSQSLL HSNGNTYLHW YLQKPGQSPK LLIYKVSNRF SGVPDRFSGS GSGTDFTLKI SRVEAEDLG VYFCSQSTHI PPWTFGGGTK LDIKRADAAP TVSIFPPSSE QLTSGGLSVV CFLNNFYPKD INVKWKIDGS E RQNGVLNS WTDQDSKDST YSMSSTLTLT KDEYERHNSY TCEATHKTST SPIVKSFNRN EC |
-Macromolecule #3: UIC2 Fab-fragment heavy chain
| Macromolecule | Name: UIC2 Fab-fragment heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 24.381281 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLQESGPE LVKTGASVKI SCKASGYSFS NYYIHWVKQS HGKSLEWIGF ISCYNGATFY NQKFKGKATF TVDNSSSTAY MKFNSLTFE DSAVYYCARL PIQFGNFYPM DYWGQGTTVT VSSAKTTAPS VYPLAPVCGD TTGSSVTLGC LVKGYFPEPV T LTWNSGSL ...String: EVQLQESGPE LVKTGASVKI SCKASGYSFS NYYIHWVKQS HGKSLEWIGF ISCYNGATFY NQKFKGKATF TVDNSSSTAY MKFNSLTFE DSAVYYCARL PIQFGNFYPM DYWGQGTTVT VSSAKTTAPS VYPLAPVCGD TTGSSVTLGC LVKGYFPEPV T LTWNSGSL SSGVHTFPAV LQSDLYTLSS SVTVTSSTWP SQSITCNVAH PASSTKVDKK IEPRGPT |
-Macromolecule #4: ~{N}-[2-[2-[4-[2-(6,7-dimethoxy-3,4-dihydro-1~{H}-isoquinolin-2-y...
| Macromolecule | Name: ~{N}-[2-[2-[4-[2-(6,7-dimethoxy-3,4-dihydro-1~{H}-isoquinolin-2-yl)ethyl]phenyl]-1,2,3,4-tetrazol-5-yl]-4,5-dimethoxy-phenyl]-4-oxidanylidene-2,3-dihydrochromene-2-carboxamide type: ligand / ID: 4 / Number of copies: 2 / Formula: V5Q |
|---|---|
| Molecular weight | Theoretical: 690.744 Da |
| Chemical component information | 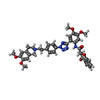 ChemComp-V5Q: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 58.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller