+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12516 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | B-brick bare in 5 mM Mg2+ | |||||||||||||||
 Map data Map data | Composite map from Multibody and PostProcessing job | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | DNA Origami / DNA | |||||||||||||||
| Biological species | synthetic construct (others) | |||||||||||||||
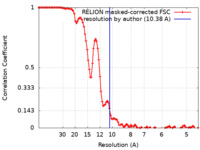
| Method | single particle reconstruction / cryo EM / Resolution: 10.38 Å | |||||||||||||||
 Authors Authors | Bertosin E / Stoemmer P | |||||||||||||||
| Funding support |  Germany, 4 items Germany, 4 items
| |||||||||||||||
 Citation Citation |  Journal: ACS Nano / Year: 2021 Journal: ACS Nano / Year: 2021Title: Cryo-Electron Microscopy and Mass Analysis of Oligolysine-Coated DNA Nanostructures. Authors: Eva Bertosin / Pierre Stömmer / Elija Feigl / Maximilian Wenig / Maximilian N Honemann / Hendrik Dietz /  Abstract: Cationic coatings can enhance the stability of synthetic DNA objects in low ionic strength environments such as physiological fluids. Here, we used single-particle cryo-electron microscopy (cryo-EM), ...Cationic coatings can enhance the stability of synthetic DNA objects in low ionic strength environments such as physiological fluids. Here, we used single-particle cryo-electron microscopy (cryo-EM), pseudoatomic model fitting, and single-molecule mass photometry to study oligolysine and polyethylene glycol (PEG)-oligolysine-coated multilayer DNA origami objects. The coatings preserve coarse structural features well on a resolution of multiple nanometers but can also induce deformations such as twisting and bending. Higher-density coatings also led to internal structural deformations in the DNA origami test objects, in which a designed honeycomb-type helical lattice was deformed into a more square-lattice-like pattern. Under physiological ionic strength, where the uncoated objects disassembled, the coated objects remained intact but they shrunk in the helical direction and expanded in the direction perpendicular to the helical axis. Helical details like major/minor grooves and crossover locations were not discernible in cryo-EM maps that we determined of DNA origami coated with oligolysine and PEG-oligolysine, whereas these features were visible in cryo-EM maps determined from the uncoated reference objects. Blunt-ended double-helical interfaces remained accessible underneath the coating and may be used for the formation of multimeric DNA origami assemblies that rely on stacking interactions between blunt-ended helices. The ionic strength requirements for forming multimers from coated DNA origami differed from those needed for uncoated objects. Using single-molecule mass photometry, we found that the mass of coated DNA origami objects prior to and after incubation in low ionic strength physiological conditions remained unchanged. This finding indicated that the coating effectively prevented strand dissociation but also that the coating itself remained stable in place. Our results validate oligolysine coatings as a powerful stabilization method for DNA origami but also reveal several potential points of failure that experimenters should watch to avoid working with false premises. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12516.map.gz emd_12516.map.gz | 7.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12516-v30.xml emd-12516-v30.xml emd-12516.xml emd-12516.xml | 93.4 KB 93.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12516_fsc.xml emd_12516_fsc.xml | 10.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_12516.png emd_12516.png | 56 KB | ||
| Filedesc metadata |  emd-12516.cif.gz emd-12516.cif.gz | 9.4 KB | ||
| Others |  emd_12516_additional_1.map.gz emd_12516_additional_1.map.gz | 5.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12516 http://ftp.pdbj.org/pub/emdb/structures/EMD-12516 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12516 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12516 | HTTPS FTP |
-Related structure data
| Related structure data |  7npnMC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12516.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12516.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Composite map from Multibody and PostProcessing job | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.319 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Refined and post-processed map
| File | emd_12516_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Refined and post-processed map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : B-brick bare in 5 mM Mg2+
+Supramolecule #1: B-brick bare in 5 mM Mg2+
+Macromolecule #1: SCAFFOLD STRAND
+Macromolecule #2: STAPLE STRAND
+Macromolecule #3: STAPLE STRAND
+Macromolecule #4: STAPLE STRAND
+Macromolecule #5: STAPLE STRAND
+Macromolecule #6: STAPLE STRAND
+Macromolecule #7: STAPLE STRAND
+Macromolecule #8: STAPLE STRAND
+Macromolecule #9: STAPLE STRAND
+Macromolecule #10: STAPLE STRAND
+Macromolecule #11: STAPLE STRAND
+Macromolecule #12: STAPLE STRAND
+Macromolecule #13: STAPLE STRAND
+Macromolecule #14: STAPLE STRAND
+Macromolecule #15: STAPLE STRAND
+Macromolecule #16: STAPLE STRAND
+Macromolecule #17: STAPLE STRAND
+Macromolecule #18: STAPLE STRAND
+Macromolecule #19: STAPLE STRAND
+Macromolecule #20: STAPLE STRAND
+Macromolecule #21: STAPLE STRAND
+Macromolecule #22: STAPLE STRAND
+Macromolecule #23: STAPLE STRAND
+Macromolecule #24: STAPLE STRAND
+Macromolecule #25: STAPLE STRAND
+Macromolecule #26: STAPLE STRAND
+Macromolecule #27: STAPLE STRAND
+Macromolecule #28: STAPLE STRAND
+Macromolecule #29: STAPLE STRAND
+Macromolecule #30: STAPLE STRAND
+Macromolecule #31: STAPLE STRAND
+Macromolecule #32: STAPLE STRAND
+Macromolecule #33: STAPLE STRAND
+Macromolecule #34: STAPLE STRAND
+Macromolecule #35: STAPLE STRAND
+Macromolecule #36: STAPLE STRAND
+Macromolecule #37: STAPLE STRAND
+Macromolecule #38: STAPLE STRAND
+Macromolecule #39: STAPLE STRAND
+Macromolecule #40: STAPLE STRAND
+Macromolecule #41: STAPLE STRAND
+Macromolecule #42: STAPLE STRAND
+Macromolecule #43: STAPLE STRAND
+Macromolecule #44: STAPLE STRAND
+Macromolecule #45: STAPLE STRAND
+Macromolecule #46: STAPLE STRAND
+Macromolecule #47: STAPLE STRAND
+Macromolecule #48: STAPLE STRAND
+Macromolecule #49: STAPLE STRAND
+Macromolecule #50: STAPLE STRAND
+Macromolecule #51: STAPLE STRAND
+Macromolecule #52: STAPLE STRAND
+Macromolecule #53: STAPLE STRAND
+Macromolecule #54: STAPLE STRAND
+Macromolecule #55: STAPLE STRAND
+Macromolecule #56: STAPLE STRAND
+Macromolecule #57: STAPLE STRAND
+Macromolecule #58: STAPLE STRAND
+Macromolecule #59: STAPLE STRAND
+Macromolecule #60: STAPLE STRAND
+Macromolecule #61: STAPLE STRAND
+Macromolecule #62: STAPLE STRAND
+Macromolecule #63: STAPLE STRAND
+Macromolecule #64: STAPLE STRAND
+Macromolecule #65: STAPLE STRAND
+Macromolecule #66: STAPLE STRAND
+Macromolecule #67: STAPLE STRAND
+Macromolecule #68: STAPLE STRAND
+Macromolecule #69: STAPLE STRAND
+Macromolecule #70: STAPLE STRAND
+Macromolecule #71: STAPLE STRAND
+Macromolecule #72: STAPLE STRAND
+Macromolecule #73: STAPLE STRAND
+Macromolecule #74: STAPLE STRAND
+Macromolecule #75: STAPLE STRAND
+Macromolecule #76: STAPLE STRAND
+Macromolecule #77: STAPLE STRAND
+Macromolecule #78: STAPLE STRAND
+Macromolecule #79: STAPLE STRAND
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)