[English] 日本語
 Yorodumi
Yorodumi- EMDB-1234: West Nile virus in complex with the Fab fragment of a neutralizin... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1234 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | West Nile virus in complex with the Fab fragment of a neutralizing monoclonal antibody. | |||||||||
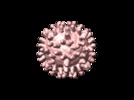
 Map data Map data | map of West Nile Virus complexed with Fab fragment of neutralizing antibody E16 | |||||||||
 Sample Sample |
| |||||||||
| Biological species | unidentified (others) /  West Nile virus West Nile virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 14.5 Å | |||||||||
 Authors Authors | Kaufmann B / Nybakken GE / Chipman PR / Zhang W / Diamond MS / Fremont DH / Kuhn RJ / Rossmann MG | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2006 Journal: Proc Natl Acad Sci U S A / Year: 2006Title: West Nile virus in complex with the Fab fragment of a neutralizing monoclonal antibody. Authors: Bärbel Kaufmann / Grant E Nybakken / Paul R Chipman / Wei Zhang / Michael S Diamond / Daved H Fremont / Richard J Kuhn / Michael G Rossmann /  Abstract: Flaviviruses, such as West Nile virus (WNV), are significant human pathogens. The humoral immune response plays an important role in the control of flavivirus infection and disease. The structure of ...Flaviviruses, such as West Nile virus (WNV), are significant human pathogens. The humoral immune response plays an important role in the control of flavivirus infection and disease. The structure of WNV complexed with the Fab fragment of the strongly neutralizing mAb E16 was determined to 14.5-Angstrom resolution with cryo-electron microscopy. E16, an antibody with therapeutic potential, binds to domain III of the WNV envelope glycoprotein. Because of steric hindrance, Fab E16 binds to only 120 of the 180 possible binding sites on the viral surface. Fitting of the previously determined x-ray structure of the Fab-domain III complex into the cryo-electron microscopy density required a change of the elbow angle between the variable and constant domains of the Fab. The structure suggests that the E16 antibody neutralizes WNV by blocking the initial rearrangement of the E glycoprotein before fusion with a cellular membrane. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1234.map.gz emd_1234.map.gz | 32.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1234-v30.xml emd-1234-v30.xml emd-1234.xml emd-1234.xml | 12 KB 12 KB | Display Display |  EMDB header EMDB header |
| Images |  1234.gif 1234.gif | 12.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1234 http://ftp.pdbj.org/pub/emdb/structures/EMD-1234 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1234 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1234 | HTTPS FTP |
-Validation report
| Summary document |  emd_1234_validation.pdf.gz emd_1234_validation.pdf.gz | 242.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1234_full_validation.pdf.gz emd_1234_full_validation.pdf.gz | 241.2 KB | Display | |
| Data in XML |  emd_1234_validation.xml.gz emd_1234_validation.xml.gz | 7.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1234 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1234 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1234 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1234 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1234.map.gz / Format: CCP4 / Size: 101.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1234.map.gz / Format: CCP4 / Size: 101.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | map of West Nile Virus complexed with Fab fragment of neutralizing antibody E16 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.7459 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
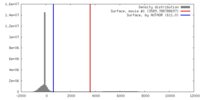
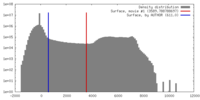
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : West Nile virus NY99 complexed with Fab fragment of neutralizing ...
| Entire | Name: West Nile virus NY99 complexed with Fab fragment of neutralizing monoclonal antibody E16 |
|---|---|
| Components |
|
-Supramolecule #1000: West Nile virus NY99 complexed with Fab fragment of neutralizing ...
| Supramolecule | Name: West Nile virus NY99 complexed with Fab fragment of neutralizing monoclonal antibody E16 type: sample / ID: 1000 Oligomeric state: T1 icosahedron with three E monomers and two Fab per asymmetric unit Number unique components: 2 |
|---|
-Supramolecule #1: West Nile virus
| Supramolecule | Name: West Nile virus / type: virus / ID: 1 / Name.synonym: WNV / NCBI-ID: 11082 / Sci species name: West Nile virus / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No / Syn species name: WNV |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
| Virus shell | Shell ID: 1 / Name: E glycoprotein / Diameter: 500 Å / T number (triangulation number): 1 |
-Macromolecule #1: E16 Fab fragment
| Macromolecule | Name: E16 Fab fragment / type: protein_or_peptide / ID: 1 / Number of copies: 120 / Oligomeric state: Monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism: unidentified (others) |
| Recombinant expression | Organism: mammalian cell line (unknown) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 / Details: 12 mM Tris-HCl, 120 mM NaCl, 1 mM EDTA |
|---|---|
| Grid | Details: 400 mesh copper grid |
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER Details: Vitrification instrument: Guillotine-style plunge freezeing device Method: A small vial of ethane is placed inside a larger liquid nitrogen reservoir. The grid holding a few microliters of the sample is held in place at the bottom of a plunger by the means of fine ...Method: A small vial of ethane is placed inside a larger liquid nitrogen reservoir. The grid holding a few microliters of the sample is held in place at the bottom of a plunger by the means of fine tweezers. Once the ethane in the vial is completely frozen, it needs to be slightly melted. When the liquid ethane is ready, a piece of filter paper is then pressed against the sample to blot of excess buffer, sufficient to leave a thin layer on the grid. After a predetermined time, the filter paper is removed, and the plunger is allowed to drop into the liquid ethane. Once the grid enters the liquid ethane, the sample is rapidly frozen, and the grid is transferred under liquid nitrogen to a storage box immersed liquid nitrogen for later use in the microscope. |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM300FEG/T |
|---|---|
| Temperature | Average: 98 K |
| Alignment procedure | Legacy - Astigmatism: live FFT at 200K mag |
| Details | low dose |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: OTHER / Digitization - Sampling interval: 6.35 µm / Number real images: 78 / Average electron dose: 22 e/Å2 / Details: Nikon SuperCoolScan 9K scanned images binned 2x2 / Bits/pixel: 16 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 47190 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.88 µm / Nominal defocus min: 1.32 µm / Nominal magnification: 47000 |
| Sample stage | Specimen holder: EUCENTRIC / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Details | The particles were selected interactively at the computer terminal. |
|---|---|
| CTF correction | Details: Each particle |
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 14.5 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: PFTSEARCH, PO2R, P3DR Details: final map includes data to 12.8 Ang resolution (fsc 0.3 cut-off); magnification of final map standardized to a map calculated from dengue virus model coordinates (PDB accession no 1THD) Number images used: 3567 |
| Final angle assignment | Details: theta 69 to 90 degrees phi -31 to 31 degrees omega 0 to 360 degrees |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)