+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12169 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | S-layer Deinoxanthin Binding Complex (SDBC) | ||||||||||||
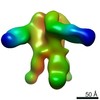
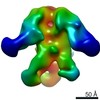
 Map data Map data | 3D cryo-EM map of the S-layer Deinoxanthin Binding Complex (SDBC) isolated from the radio-resistant bacterium Deinococcus radiodurans | ||||||||||||
 Sample Sample |
| ||||||||||||
| Biological species |  Deinococcus radiodurans R1 (radioresistant) Deinococcus radiodurans R1 (radioresistant) | ||||||||||||
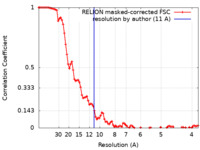
| Method | single particle reconstruction / cryo EM / Resolution: 11.0 Å | ||||||||||||
 Authors Authors | Farci D / Kereiche S / Piano D | ||||||||||||
| Funding support |  Poland, Poland,  Czech Republic, 3 items Czech Republic, 3 items
| ||||||||||||
 Citation Citation | Journal: J Biol Chem / Year: 2020 Title: Structural insights into the main S-layer unit of reveal a massive protein complex with porin-like features. Authors: Domenica Farci / Mehmet Alphan Aksoyoglu / Stefano Francesco Farci / Jayesh Arun Bafna / Igor Bodrenko / Matteo Ceccarelli / Joanna Kirkpatrick / Mathias Winterhalter / Sami Kereïche / Dario Piano /      Abstract: In the extremophile bacterium , the outermost surface layer is tightly connected with the rest of the cell wall. This integrated organization provides a compact structure that shields the bacterium ...In the extremophile bacterium , the outermost surface layer is tightly connected with the rest of the cell wall. This integrated organization provides a compact structure that shields the bacterium against environmental stresses. The fundamental unit of this surface layer (S-layer) is the S-layer deinoxanthin-binding complex (SDBC), which binds the carotenoid deinoxanthin and provides both, thermostability and UV radiation resistance. However, the structural organization of the SDBC awaits elucidation. Here, we report the isolation of the SDBC with a gentle procedure consisting of lysozyme treatment and solubilization with the nonionic detergent -dodecyl-β-d-maltoside, which preserved both hydrophilic and hydrophobic components of the SDBC and allows the retention of several minor subunits. As observed by low-resolution single-particle analysis, we show that the complex possesses a porin-like structural organization, but is larger than other known porins. We also noted that the main SDBC component, the protein DR_2577, shares regions of similarity with known porins. Moreover, results from electrophysiological assays with membrane-reconstituted SDBC disclosed that it is a nonselective channel that has some peculiar gating properties, but also exhibits behavior typically observed in pore-forming proteins, such as porins and ionic transporters. The functional properties of this system and its porin-like organization provide information critical for understanding ion permeability through the outer cell surface of S-layer-carrying bacterial species. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12169.map.gz emd_12169.map.gz | 59.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12169-v30.xml emd-12169-v30.xml emd-12169.xml emd-12169.xml | 16.6 KB 16.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12169_fsc.xml emd_12169_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_12169.png emd_12169.png | 66.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12169 http://ftp.pdbj.org/pub/emdb/structures/EMD-12169 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12169 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12169 | HTTPS FTP |
-Validation report
| Summary document |  emd_12169_validation.pdf.gz emd_12169_validation.pdf.gz | 417.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_12169_full_validation.pdf.gz emd_12169_full_validation.pdf.gz | 416.6 KB | Display | |
| Data in XML |  emd_12169_validation.xml.gz emd_12169_validation.xml.gz | 10.7 KB | Display | |
| Data in CIF |  emd_12169_validation.cif.gz emd_12169_validation.cif.gz | 14.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12169 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12169 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12169 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12169 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_12169.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12169.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D cryo-EM map of the S-layer Deinoxanthin Binding Complex (SDBC) isolated from the radio-resistant bacterium Deinococcus radiodurans | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.92 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
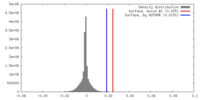
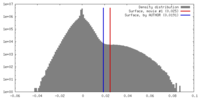
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : S-layer Deinoxanthin Binding Complex (SDBC)
| Entire | Name: S-layer Deinoxanthin Binding Complex (SDBC) |
|---|---|
| Components |
|
-Supramolecule #1: S-layer Deinoxanthin Binding Complex (SDBC)
| Supramolecule | Name: S-layer Deinoxanthin Binding Complex (SDBC) / type: complex / ID: 1 / Parent: 0 Details: The SDBC is a hetero-oligomeric complex that constitutes the main protein complex of the S-layer of the radio-resistant bacterium Deinococcus radiodurans. This complex provides protection ...Details: The SDBC is a hetero-oligomeric complex that constitutes the main protein complex of the S-layer of the radio-resistant bacterium Deinococcus radiodurans. This complex provides protection against UV radiation and shows gating properties by allowing ions currents across the cell wall. |
|---|---|
| Source (natural) | Organism:  Deinococcus radiodurans R1 (radioresistant) Deinococcus radiodurans R1 (radioresistant) |
| Molecular weight | Experimental: 800 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R2/1 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Number real images: 1573 / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 2.7 µm / Calibrated defocus max: 3.3000000000000003 µm / Calibrated defocus min: 1.0 µm / Calibrated magnification: 59000 / Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal magnification: 59000 |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)