+ Open data
Open data
- Basic information
Basic information
| Entry | Database: SASBDB / ID: SASDBR9 |
|---|---|
 Sample Sample | Human fibroblast growth factor receptor 3, from SEC-SAXS
|
| Function / homology |  Function and homology information Function and homology informationfibroblast growth factor receptor apoptotic signaling pathway / t(4;14) translocations of FGFR3 / negative regulation of developmental growth / Signaling by FGFR3 fusions in cancer /  bone maturation / chondrocyte proliferation / endochondral bone growth / FGFR3b ligand binding and activation / Signaling by activated point mutants of FGFR3 / FGFR3c ligand binding and activation ...fibroblast growth factor receptor apoptotic signaling pathway / t(4;14) translocations of FGFR3 / negative regulation of developmental growth / Signaling by FGFR3 fusions in cancer / bone maturation / chondrocyte proliferation / endochondral bone growth / FGFR3b ligand binding and activation / Signaling by activated point mutants of FGFR3 / FGFR3c ligand binding and activation ...fibroblast growth factor receptor apoptotic signaling pathway / t(4;14) translocations of FGFR3 / negative regulation of developmental growth / Signaling by FGFR3 fusions in cancer /  bone maturation / chondrocyte proliferation / endochondral bone growth / FGFR3b ligand binding and activation / Signaling by activated point mutants of FGFR3 / FGFR3c ligand binding and activation / Phospholipase C-mediated cascade; FGFR3 / bone maturation / chondrocyte proliferation / endochondral bone growth / FGFR3b ligand binding and activation / Signaling by activated point mutants of FGFR3 / FGFR3c ligand binding and activation / Phospholipase C-mediated cascade; FGFR3 /  fibroblast growth factor receptor activity / fibroblast growth factor receptor activity /  endochondral ossification / positive regulation of phospholipase activity / bone morphogenesis / PI-3K cascade:FGFR3 / endochondral ossification / positive regulation of phospholipase activity / bone morphogenesis / PI-3K cascade:FGFR3 /  fibroblast growth factor binding / fibroblast growth factor binding /  bone mineralization / cell surface receptor signaling pathway via JAK-STAT / PI3K Cascade / fibroblast growth factor receptor signaling pathway / chondrocyte differentiation / SHC-mediated cascade:FGFR3 / FRS-mediated FGFR3 signaling / positive regulation of tyrosine phosphorylation of STAT protein / bone mineralization / cell surface receptor signaling pathway via JAK-STAT / PI3K Cascade / fibroblast growth factor receptor signaling pathway / chondrocyte differentiation / SHC-mediated cascade:FGFR3 / FRS-mediated FGFR3 signaling / positive regulation of tyrosine phosphorylation of STAT protein /  transport vesicle / Signaling by FGFR3 in disease / transport vesicle / Signaling by FGFR3 in disease /  skeletal system development / Negative regulation of FGFR3 signaling / skeletal system development / Negative regulation of FGFR3 signaling /  cell surface receptor protein tyrosine kinase signaling pathway / cell surface receptor protein tyrosine kinase signaling pathway /  receptor protein-tyrosine kinase / Constitutive Signaling by Aberrant PI3K in Cancer / receptor protein-tyrosine kinase / Constitutive Signaling by Aberrant PI3K in Cancer /  MAPK cascade / PIP3 activates AKT signaling / cell-cell signaling / PI5P, PP2A and IER3 Regulate PI3K/AKT Signaling / RAF/MAP kinase cascade / MAPK cascade / PIP3 activates AKT signaling / cell-cell signaling / PI5P, PP2A and IER3 Regulate PI3K/AKT Signaling / RAF/MAP kinase cascade /  protein tyrosine kinase activity / positive regulation of MAPK cascade / positive regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / positive regulation of ERK1 and ERK2 cascade / protein tyrosine kinase activity / positive regulation of MAPK cascade / positive regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / positive regulation of ERK1 and ERK2 cascade /  receptor complex / receptor complex /  phosphorylation / positive regulation of cell population proliferation / phosphorylation / positive regulation of cell population proliferation /  Golgi apparatus / Golgi apparatus /  cell surface / cell surface /  endoplasmic reticulum / extracellular region / endoplasmic reticulum / extracellular region /  ATP binding / identical protein binding / ATP binding / identical protein binding /  plasma membrane plasma membraneSimilarity search - Function |
| Biological species |   Homo sapiens (human) Homo sapiens (human) |
 Citation Citation |  Date: 2018 Mar Date: 2018 MarTitle: Disease Variants of FGFR3 Reveal Molecular Basis for the Recognition and Additional Roles for Cdc37 in Hsp90 Chaperone System Authors: Bunney T / Inglis A / Sanfelice D / Farrell B / Kerr C / Thompson G / Masson G / Thiyagarajan N / Svergun D / Williams R / Breeze A |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDBR9 SASDBR9 |
|---|
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- External links
External links
| Related items in Molecule of the Month |
|---|
-Models
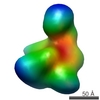
| Model #1134 |  Type: mix / Software: (ATSAS2.7.2) / Radius of dummy atoms: 3.50 A / Symmetry  : P1 / Chi-square value: 3.10 : P1 / Chi-square value: 3.10 Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
|---|---|
| Model #1135 |  Type: dummy / Software: (ATSAS2.7.2) / Radius of dummy atoms: 3.00 A / Symmetry  : P1 / Chi-square value: 1.201 : P1 / Chi-square value: 1.201 Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
- Sample
Sample
 Sample Sample | Name: Human fibroblast growth factor receptor 3, from SEC-SAXS Specimen concentration: 2 mg/ml |
|---|---|
| Buffer | Name: 25 mM Tris.Cl, 150 mM NaCl, 5% (v/v) glycerol, 1 mM TCEP pH: 8 |
| Entity #589 | Name: FGFR3 Fibroblast growth factor receptor 3 / Type: protein / Description: Fibroblast growth factor receptor 3 Fibroblast growth factor receptor 3 / Type: protein / Description: Fibroblast growth factor receptor 3 / Formula weight: 35.478 / Num. of mol.: 1 / Source: Homo sapiens / References: UniProt: P22607 / Formula weight: 35.478 / Num. of mol.: 1 / Source: Homo sapiens / References: UniProt: P22607Sequence: SELELPADPK WELSRARLTL GKPLGEGAFG QVVMAEAIGI DKDRAAKPVT VAVKMLKDDA TDKDLSDLVS EMEMMKMIGK HKNFINLLGA CTQGGPLYVL VEYAAKGNLR EFLRARRPPG LDYSFDTSKP PEEQLTFKDL VSCAYQVARG MEYLASQKCI HRDLAARNVL ...Sequence: SELELPADPK WELSRARLTL GKPLGEGAFG QVVMAEAIGI DKDRAAKPVT VAVKMLKDDA TDKDLSDLVS EMEMMKMIGK HKNFINLLGA CTQGGPLYVL VEYAAKGNLR EFLRARRPPG LDYSFDTSKP PEEQLTFKDL VSCAYQVARG MEYLASQKCI HRDLAARNVL VTEDNVMKIA DFGLARDVHN LDYYKKTTNG RLPVKWMAPE ALFDRVYTHQ SDVWSFGVLL WEIFTLGGSP YPGIPVEELF KLLKEGHRMD KPANCTHDLY MIMRECWHAA PSQRPTFKQL VEDLDRVLTV TSTDEYLDLS APFE |
-Experimental information
| Beam | Instrument name: PETRA III P12 / City: Hamburg / 国: Germany  / Type of source: X-ray synchrotron / Type of source: X-ray synchrotron Synchrotron / Wavelength: 0.124 Å / Dist. spec. to detc.: 3.1 mm Synchrotron / Wavelength: 0.124 Å / Dist. spec. to detc.: 3.1 mm | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Pilatus 2M | ||||||||||||||||||||||||||||||||||||
| Scan | Measurement date: Jul 10, 2016 / Storage temperature: 20 °C / Cell temperature: 20 °C / Exposure time: 1 sec. / Number of frames: 3600 / Unit: 1/nm /
| ||||||||||||||||||||||||||||||||||||
| Distance distribution function P(R) |
| ||||||||||||||||||||||||||||||||||||
| Result |
|
 Movie
Movie Controller
Controller