+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8w2r | ||||||
|---|---|---|---|---|---|---|---|
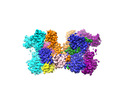
| Title | HIV-1 P5-IN intasome core | ||||||
 Components Components |
| ||||||
 Keywords Keywords |  VIRAL PROTEIN / VIRAL PROTEIN /  HIV-1 / HIV-1 /  integrase / nucleoprotein complexes / INSTI / intasome / integrase / nucleoprotein complexes / INSTI / intasome /  CryoEM CryoEM | ||||||
| Function / homology |  Function and homology information Function and homology informationDNA integration / viral genome integration into host DNA / establishment of integrated proviral latency /  RNA-directed DNA polymerase activity / RNA-directed DNA polymerase activity /  endonuclease activity / DNA recombination / symbiont entry into host cell / endonuclease activity / DNA recombination / symbiont entry into host cell /  DNA binding / zinc ion binding DNA binding / zinc ion bindingSimilarity search - Function | ||||||
| Biological species |    Human immunodeficiency virus 1 Human immunodeficiency virus 1synthetic construct (others) | ||||||
| Method |  ELECTRON MICROSCOPY / ELECTRON MICROSCOPY /  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.23 Å cryo EM / Resolution: 3.23 Å | ||||||
 Authors Authors | Li, M. / Craigie, R. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2024 Journal: J Mol Biol / Year: 2024Title: HIV-1 Integrase Assembles Multiple Species of Stable Synaptic Complex Intasomes That Are Active for Concerted DNA Integration In vitro. Authors: Min Li / Renbin Yang / Xuemin Chen / Huaibin Wang / Rodolfo Ghirlando / Emilios K Dimitriadis / Robert Craigie /  Abstract: Retroviral DNA integration is mediated by nucleoprotein complexes (intasomes) in which a pair of viral DNA ends are bridged by a multimer of integrase (IN). Most of the high-resolution structures of ...Retroviral DNA integration is mediated by nucleoprotein complexes (intasomes) in which a pair of viral DNA ends are bridged by a multimer of integrase (IN). Most of the high-resolution structures of HIV-1 intasomes are based on an HIV-1 IN with an Sso7d protein domain fused to the N-terminus. Sso7d-IN aggregates much less than wild-type IN and has been critical for structural studies of HIV-1 intasomes. Unexpectedly, these structures revealed that the common core architecture that mediates catalysis could be assembled in various ways, giving rise to both tetrameric and dodecameric intasomes, together with other less well-characterized species. This differs from related retroviruses that assemble unique multimeric intasomes, although the number of protomers in the intasome varies between viruses. The question of whether the additional Sso7d domain contributes to the heterogeneity of HIV-1 intasomes is therefore raised. We have addressed this by biochemical and structural studies of intasomes assembled with wild-type HIV-1 IN. Negative stain and cryo-EM reveal a similar range of multimeric intasome species as with Sso7d-IN with the same common core architecture. Stacks of intasomes resulting from domain swapping are also seen with both wild-type and Sso7d-IN intasomes. The propensity to assemble multimeric intasome species is, therefore, an intrinsic property of HIV-1 IN and is not conferred by the presence of the Sso7d domain. The recently solved intasome structures of different retroviral species, which have been reported to be tetrameric, octameric, dodecameric, and hexadecameric, highlight how a common intasome core architecture can be assembled in different ways for catalysis. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8w2r.cif.gz 8w2r.cif.gz | 340.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8w2r.ent.gz pdb8w2r.ent.gz | 236.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8w2r.json.gz 8w2r.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/w2/8w2r https://data.pdbj.org/pub/pdb/validation_reports/w2/8w2r ftp://data.pdbj.org/pub/pdb/validation_reports/w2/8w2r ftp://data.pdbj.org/pub/pdb/validation_reports/w2/8w2r | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  43756MC  8w09C  8w34C M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
-Protein , 1 types, 8 molecules ABCDIKLM
| #1: Protein |  Mass: 39850.418 Da / Num. of mol.: 8 Source method: isolated from a genetically manipulated source Source: (gene. exp.)    Human immunodeficiency virus 1 / Gene: pol / Production host: Human immunodeficiency virus 1 / Gene: pol / Production host:   Escherichia coli (E. coli) / References: UniProt: F2WR39 Escherichia coli (E. coli) / References: UniProt: F2WR39 |
|---|
-DNA chain , 2 types, 4 molecules ENFO
| #2: DNA chain | Mass: 8188.271 Da / Num. of mol.: 2 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) #3: DNA chain | Mass: 7773.023 Da / Num. of mol.: 2 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) |
|---|
-Non-polymers , 3 types, 8 molecules 



| #4: Chemical | ChemComp-MG / #5: Chemical | #6: Chemical |  Dolutegravir Dolutegravir |
|---|
-Details
| Has ligand of interest | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method:  single particle reconstruction single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: HIV-1 P5-IN intasomes Core / Type: COMPLEX / Details: HIV-1 intasomes assembled with P5-IN / Entity ID: #1-#3 / Source: RECOMBINANT |
|---|---|
| Molecular weight | Experimental value: NO |
| Source (natural) | Organism:    Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Source (recombinant) | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Buffer solution | pH: 6.2 |
| Specimen | Conc.: 0.4 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied : NO / Vitrification applied : NO / Vitrification applied : YES : YES |
Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source : :  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy / Nominal magnification: 105000 X / Nominal defocus max: 2600 nm / Nominal defocus min: 1400 nm / Cs Bright-field microscopy / Nominal magnification: 105000 X / Nominal defocus max: 2600 nm / Nominal defocus min: 1400 nm / Cs : 2.7 mm / C2 aperture diameter: 70 µm / Alignment procedure: COMA FREE : 2.7 mm / C2 aperture diameter: 70 µm / Alignment procedure: COMA FREE |
| Specimen holder | Cryogen: NITROGEN / Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Image recording | Electron dose: 54 e/Å2 / Detector mode: SUPER-RESOLUTION / Film or detector model: GATAN K2 SUMMIT (4k x 4k) |
| EM imaging optics | Energyfilter name : GIF Bioquantum / Energyfilter slit width: 20 eV : GIF Bioquantum / Energyfilter slit width: 20 eV |
- Processing
Processing
| EM software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
CTF correction | Type: NONE | ||||||||||||||||||||||||
| Particle selection | Num. of particles selected: 643000 | ||||||||||||||||||||||||
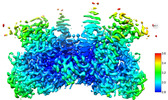
3D reconstruction | Resolution: 3.23 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 154000 / Symmetry type: POINT | ||||||||||||||||||||||||
| Atomic model building | Protocol: RIGID BODY FIT / Space: REAL | ||||||||||||||||||||||||
| Atomic model building | PDB-ID: 5U1C Accession code: 5U1C / Source name: PDB / Type: experimental model | ||||||||||||||||||||||||
| Refinement | Cross valid method: NONE |
 Movie
Movie Controller
Controller



 PDBj
PDBj











































