[English] 日本語
 Yorodumi
Yorodumi- PDB-8t56: Structure of mechanically activated ion channel OSCA1.2 in peptidiscs -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8t56 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Structure of mechanically activated ion channel OSCA1.2 in peptidiscs | ||||||
 Components Components |
| ||||||
 Keywords Keywords |  MEMBRANE PROTEIN / mechanically activated ion channel MEMBRANE PROTEIN / mechanically activated ion channel | ||||||
| Function / homology |  Function and homology information Function and homology informationmechanosensitive monoatomic ion channel activity / calcium-activated cation channel activity / monoatomic cation transport / identical protein binding /  plasma membrane plasma membraneSimilarity search - Function | ||||||
| Biological species |   Arabidopsis thaliana (thale cress) Arabidopsis thaliana (thale cress)synthetic construct (others) | ||||||
| Method |  ELECTRON MICROSCOPY / ELECTRON MICROSCOPY /  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.8 Å cryo EM / Resolution: 2.8 Å | ||||||
 Authors Authors | Burendei, B. / Jojoa-Cruz, S. / Lee, W.H. / Ward, A.B. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Structure / Year: 2024 Journal: Structure / Year: 2024Title: Structure of mechanically activated ion channel OSCA2.3 reveals mobile elements in the transmembrane domain. Authors: Sebastian Jojoa-Cruz / Batuujin Burendei / Wen-Hsin Lee / Andrew B Ward /  Abstract: Members of the OSCA/TMEM63 family are mechanically activated ion channels and structures of some OSCA members have revealed the architecture of these channels and structural features that are ...Members of the OSCA/TMEM63 family are mechanically activated ion channels and structures of some OSCA members have revealed the architecture of these channels and structural features that are potentially involved in mechanosensation. However, these structures are all in a similar state and information about the motion of different elements of the structure is limited, preventing a deeper understanding of how these channels work. Here, we used cryoelectron microscopy to determine high-resolution structures of Arabidopsis thaliana OSCA1.2 and OSCA2.3 in peptidiscs. The structure of OSCA1.2 matches previous structures of the same protein in different environments. Yet, in OSCA2.3, the TM6a-TM7 linker adopts a different conformation that constricts the pore on its cytoplasmic side. Furthermore, coevolutionary sequence analysis uncovered a conserved interaction between the TM6a-TM7 linker and the beam-like domain (BLD). Our results reveal conformational heterogeneity and differences in conserved interactions between the TMD and BLD among members of the OSCA family. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8t56.cif.gz 8t56.cif.gz | 328.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8t56.ent.gz pdb8t56.ent.gz | 266.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8t56.json.gz 8t56.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/t5/8t56 https://data.pdbj.org/pub/pdb/validation_reports/t5/8t56 ftp://data.pdbj.org/pub/pdb/validation_reports/t5/8t56 ftp://data.pdbj.org/pub/pdb/validation_reports/t5/8t56 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 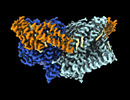 41043MC  8t57C M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
-Protein / Protein/peptide , 2 types, 10 molecules ABCDEFGHIJ
| #1: Protein | Mass: 89031.719 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Details: The last 10 residues (GTGTLEVLFQ) are leftover of a linker and protease site. Source: (gene. exp.)   Arabidopsis thaliana (thale cress) / Gene: CSC1, At4g22120, F1N20.220 / Plasmid: pcDNA3.1 / Cell line (production host): HEK293F / Production host: Arabidopsis thaliana (thale cress) / Gene: CSC1, At4g22120, F1N20.220 / Plasmid: pcDNA3.1 / Cell line (production host): HEK293F / Production host:   Homo sapiens (human) / References: UniProt: Q5XEZ5 Homo sapiens (human) / References: UniProt: Q5XEZ5#2: Protein/peptide | Mass: 4482.132 Da / Num. of mol.: 8 / Source method: obtained synthetically / Details: Synthesized by Peptidisc Biotech / Source: (synth.) synthetic construct (others) |
|---|
-Non-polymers , 4 types, 36 molecules 






| #3: Chemical | | #4: Chemical | ChemComp-PLM /  Palmitic acid Palmitic acid#5: Chemical |  POPC POPC#6: Chemical | ChemComp-CLR /  Cholesterol Cholesterol |
|---|
-Details
| Has ligand of interest | N |
|---|
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method:  single particle reconstruction single particle reconstruction |
- Sample preparation
Sample preparation
| Component |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular weight | Value: 0.176 MDa / Experimental value: NO | ||||||||||||||||||||||||
| Source (natural) |
| ||||||||||||||||||||||||
| Source (recombinant) | Organism:   Homo sapiens (human) / Cell: HEK293F / Plasmid Homo sapiens (human) / Cell: HEK293F / Plasmid : pcDNA3.1 : pcDNA3.1 | ||||||||||||||||||||||||
| Buffer solution | pH: 8 | ||||||||||||||||||||||||
| Buffer component |
| ||||||||||||||||||||||||
| Specimen | Conc.: 8 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied : NO / Vitrification applied : NO / Vitrification applied : YES : YES | ||||||||||||||||||||||||
| Specimen support | Grid material: GOLD / Grid mesh size: 300 divisions/in. | ||||||||||||||||||||||||
Vitrification | Instrument: FEI VITROBOT MARK IV / Cryogen name: ETHANE / Humidity: 100 % / Chamber temperature: 277 K |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source : :  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy / Nominal magnification: 29000 X / Nominal defocus max: 1400 nm / Nominal defocus min: 600 nm / Cs Bright-field microscopy / Nominal magnification: 29000 X / Nominal defocus max: 1400 nm / Nominal defocus min: 600 nm / Cs : 2.7 mm / C2 aperture diameter: 70 µm / Alignment procedure: COMA FREE : 2.7 mm / C2 aperture diameter: 70 µm / Alignment procedure: COMA FREE |
| Specimen holder | Cryogen: NITROGEN / Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Image recording | Electron dose: 50 e/Å2 / Detector mode: COUNTING / Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Num. of grids imaged: 1 / Num. of real images: 1805 |
| Image scans | Width: 3710 / Height: 3838 / Movie frames/image: 36 |
- Processing
Processing
| EM software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Particle selection | Num. of particles selected: 1046049 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Point symmetry : C2 (2 fold cyclic : C2 (2 fold cyclic ) ) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
3D reconstruction | Resolution: 2.8 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 52250 / Details: SIDESPLITTER was used for reconstruction step / Symmetry type: POINT | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Atomic model building | Protocol: RIGID BODY FIT / Space: REAL | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Atomic model building | PDB-ID: 6MGV Accession code: 6MGV / Source name: PDB / Type: experimental model |
 Movie
Movie Controller
Controller






 PDBj
PDBj











