+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8gbj | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of a human BCDX2/ssDNA complex | ||||||||||||||||||||||||||||||
 Components Components |
| ||||||||||||||||||||||||||||||
 Keywords Keywords |  RECOMBINATION / RECOMBINATION /  DNA binding protein / ATP binding domain / DNA binding protein / ATP binding domain /  homologous recombination / RAD51 paralog / homologous recombination / RAD51 paralog /  ssDNA ssDNA | ||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationmeiotic DNA recombinase assembly / Rad51C-XRCC3 complex / Rad51B-Rad51C-Rad51D-XRCC2 complex / female meiosis sister chromatid cohesion / blastocyst growth / crossover junction DNA endonuclease activity / somite development / DNA strand invasion / telomere maintenance via recombination / Impaired BRCA2 binding to PALB2 ...meiotic DNA recombinase assembly / Rad51C-XRCC3 complex / Rad51B-Rad51C-Rad51D-XRCC2 complex / female meiosis sister chromatid cohesion / blastocyst growth / crossover junction DNA endonuclease activity / somite development / DNA strand invasion / telomere maintenance via recombination / Impaired BRCA2 binding to PALB2 / gamma-tubulin binding / reciprocal meiotic recombination / regulation of fibroblast apoptotic process / sister chromatid cohesion /  centrosome cycle / Defective homologous recombination repair (HRR) due to BRCA1 loss of function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA1 binding function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA2/RAD51/RAD51C binding function / Homologous DNA Pairing and Strand Exchange / Resolution of D-loop Structures through Synthesis-Dependent Strand Annealing (SDSA) / Resolution of D-loop Structures through Holliday Junction Intermediates / positive regulation of neurogenesis / ATP-dependent DNA damage sensor activity / male meiosis I / Presynaptic phase of homologous DNA pairing and strand exchange / response to X-ray / ATP-dependent activity, acting on DNA / interstrand cross-link repair / centrosome cycle / Defective homologous recombination repair (HRR) due to BRCA1 loss of function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA1 binding function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA2/RAD51/RAD51C binding function / Homologous DNA Pairing and Strand Exchange / Resolution of D-loop Structures through Synthesis-Dependent Strand Annealing (SDSA) / Resolution of D-loop Structures through Holliday Junction Intermediates / positive regulation of neurogenesis / ATP-dependent DNA damage sensor activity / male meiosis I / Presynaptic phase of homologous DNA pairing and strand exchange / response to X-ray / ATP-dependent activity, acting on DNA / interstrand cross-link repair /  somitogenesis / positive regulation of G2/M transition of mitotic cell cycle / somitogenesis / positive regulation of G2/M transition of mitotic cell cycle /  telomere maintenance / telomere maintenance /  neurogenesis / neurogenesis /  replication fork / meiotic cell cycle / response to gamma radiation / TP53 Regulates Transcription of DNA Repair Genes / double-strand break repair via homologous recombination / multicellular organism growth / HDR through Homologous Recombination (HRR) / replication fork / meiotic cell cycle / response to gamma radiation / TP53 Regulates Transcription of DNA Repair Genes / double-strand break repair via homologous recombination / multicellular organism growth / HDR through Homologous Recombination (HRR) /  Meiotic recombination / Meiotic recombination /  single-stranded DNA binding / mitotic cell cycle / single-stranded DNA binding / mitotic cell cycle /  cell junction / Factors involved in megakaryocyte development and platelet production / cell junction / Factors involved in megakaryocyte development and platelet production /  double-stranded DNA binding / double-stranded DNA binding /  spermatogenesis / DNA recombination / in utero embryonic development / negative regulation of neuron apoptotic process / spermatogenesis / DNA recombination / in utero embryonic development / negative regulation of neuron apoptotic process /  chromosome, telomeric region / chromosome, telomeric region /  regulation of cell cycle / regulation of cell cycle /  DNA repair / intracellular membrane-bounded organelle / DNA repair / intracellular membrane-bounded organelle /  centrosome / positive regulation of cell population proliferation / perinuclear region of cytoplasm / centrosome / positive regulation of cell population proliferation / perinuclear region of cytoplasm /  ATP hydrolysis activity / ATP hydrolysis activity /  mitochondrion / mitochondrion /  DNA binding / DNA binding /  nucleoplasm / nucleoplasm /  ATP binding / ATP binding /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||||||||||||||||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human)synthetic construct (others) | ||||||||||||||||||||||||||||||
| Method |  ELECTRON MICROSCOPY / ELECTRON MICROSCOPY /  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.11 Å cryo EM / Resolution: 3.11 Å | ||||||||||||||||||||||||||||||
 Authors Authors | Jia, L. / Wasmuth, E.V. / Ruben, E.A. / Sung, P. / Rawal, Y. / Greene, E.C. / Meir, A. / Olsen, S.K. | ||||||||||||||||||||||||||||||
| Funding support |  United States, 9items United States, 9items
| ||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2023 Journal: Nature / Year: 2023Title: Structural insights into BCDX2 complex function in homologous recombination. Authors: Yashpal Rawal / Lijia Jia / Aviv Meir / Shuo Zhou / Hardeep Kaur / Eliza A Ruben / Youngho Kwon / Kara A Bernstein / Maria Jasin / Alexander B Taylor / Sandeep Burma / Robert Hromas / ...Authors: Yashpal Rawal / Lijia Jia / Aviv Meir / Shuo Zhou / Hardeep Kaur / Eliza A Ruben / Youngho Kwon / Kara A Bernstein / Maria Jasin / Alexander B Taylor / Sandeep Burma / Robert Hromas / Alexander V Mazin / Weixing Zhao / Daohong Zhou / Elizabeth V Wasmuth / Eric C Greene / Patrick Sung / Shaun K Olsen /  Abstract: Homologous recombination (HR) fulfils a pivotal role in the repair of DNA double-strand breaks and collapsed replication forks. HR depends on the products of several paralogues of RAD51, including ...Homologous recombination (HR) fulfils a pivotal role in the repair of DNA double-strand breaks and collapsed replication forks. HR depends on the products of several paralogues of RAD51, including the tetrameric complex of RAD51B, RAD51C, RAD51D and XRCC2 (BCDX2). BCDX2 functions as a mediator of nucleoprotein filament assembly by RAD51 and single-stranded DNA (ssDNA) during HR, but its mechanism remains undefined. Here we report cryogenic electron microscopy reconstructions of human BCDX2 in apo and ssDNA-bound states. The structures reveal how the amino-terminal domains of RAD51B, RAD51C and RAD51D participate in inter-subunit interactions that underpin complex formation and ssDNA-binding specificity. Single-molecule DNA curtain analysis yields insights into how BCDX2 enhances RAD51-ssDNA nucleoprotein filament assembly. Moreover, our cryogenic electron microscopy and functional analyses explain how RAD51C alterations found in patients with cancer inactivate DNA binding and the HR mediator activity of BCDX2. Our findings shed light on the role of BCDX2 in HR and provide a foundation for understanding how pathogenic alterations in BCDX2 impact genome repair. | ||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8gbj.cif.gz 8gbj.cif.gz | 201.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8gbj.ent.gz pdb8gbj.ent.gz | 155.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8gbj.json.gz 8gbj.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/gb/8gbj https://data.pdbj.org/pub/pdb/validation_reports/gb/8gbj ftp://data.pdbj.org/pub/pdb/validation_reports/gb/8gbj ftp://data.pdbj.org/pub/pdb/validation_reports/gb/8gbj | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 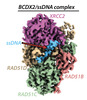 29917MC  8fazC C: citing same article ( M: map data used to model this data |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
-DNA repair protein RAD51 homolog ... , 3 types, 3 molecules BCD
| #1: Protein |  / R51H2 / RAD51 homolog B / Rad51B / RAD51-like protein 1 / R51H2 / RAD51 homolog B / Rad51B / RAD51-like protein 1Mass: 39125.855 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: RAD51B, RAD51L1, REC2 / Production host: Homo sapiens (human) / Gene: RAD51B, RAD51L1, REC2 / Production host:   Trichoplusia ni (cabbage looper) / Variant (production host): BTI-Tn-5B1-4 / References: UniProt: O15315 Trichoplusia ni (cabbage looper) / Variant (production host): BTI-Tn-5B1-4 / References: UniProt: O15315 |
|---|---|
| #2: Protein |  / R51H3 / RAD51 homolog C / RAD51-like protein 2 / R51H3 / RAD51 homolog C / RAD51-like protein 2Mass: 42244.609 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: RAD51C, RAD51L2 / Production host: Homo sapiens (human) / Gene: RAD51C, RAD51L2 / Production host:   Trichoplusia ni (cabbage looper) / Variant (production host): BTI-Tn-5B1-4 / References: UniProt: O43502 Trichoplusia ni (cabbage looper) / Variant (production host): BTI-Tn-5B1-4 / References: UniProt: O43502 |
| #3: Protein |  / R51H3 / RAD51 homolog D / RAD51-like protein 3 / TRAD / R51H3 / RAD51 homolog D / RAD51-like protein 3 / TRADMass: 35084.254 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: RAD51D, RAD51L3 / Production host: Homo sapiens (human) / Gene: RAD51D, RAD51L3 / Production host:   Trichoplusia ni (cabbage looper) / Variant (production host): BTI-Tn-5B1-4 / References: UniProt: O75771 Trichoplusia ni (cabbage looper) / Variant (production host): BTI-Tn-5B1-4 / References: UniProt: O75771 |
-Protein / DNA chain / Non-polymers , 3 types, 5 molecules XG

| #4: Protein |  / X-ray repair cross-complementing protein 2 / X-ray repair cross-complementing protein 2Mass: 32992.488 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: XRCC2 / Production host: Homo sapiens (human) / Gene: XRCC2 / Production host:   Trichoplusia ni (cabbage looper) / Variant (production host): BTI-Tn-5B1-4 / References: UniProt: O43543 Trichoplusia ni (cabbage looper) / Variant (production host): BTI-Tn-5B1-4 / References: UniProt: O43543 |
|---|---|
| #5: DNA chain | Mass: 8630.495 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) |
| #6: Chemical |
-Details
| Has ligand of interest | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method:  single particle reconstruction single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Cryo-EM structure of heterotetrameric human BCDX2 in complex with ssDNA Type: COMPLEX / Entity ID: #1-#5 / Source: RECOMBINANT |
|---|---|
| Molecular weight | Experimental value: NO |
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Source (recombinant) | Organism:   Trichoplusia ni (cabbage looper) / Strain: High Five (BTI-TN-5B1-4) Trichoplusia ni (cabbage looper) / Strain: High Five (BTI-TN-5B1-4) |
| Buffer solution | pH: 7.5 |
| Specimen | Conc.: 0.9 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied : NO / Vitrification applied : NO / Vitrification applied : YES : YES |
Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Microscopy | Model: TFS GLACIOS |
|---|---|
| Electron gun | Electron source : :  FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2000 nm / Nominal defocus min: 800 nm / Cs Bright-field microscopy / Nominal defocus max: 2000 nm / Nominal defocus min: 800 nm / Cs : 2.7 mm / C2 aperture diameter: 50 µm : 2.7 mm / C2 aperture diameter: 50 µm |
| Image recording | Electron dose: 51 e/Å2 / Film or detector model: FEI FALCON IV (4k x 4k) |
- Processing
Processing
| Software | Name: PHENIX / Version: 1.20.1_4487: / Classification: refinement | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||
3D reconstruction | Resolution: 3.11 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 165968 / Symmetry type: POINT | ||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller



 PDBj
PDBj



















































