+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7lp5 | ||||||
|---|---|---|---|---|---|---|---|
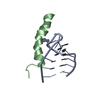
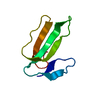
| Title | Structure of Nedd4L WW3 domain | ||||||
 Components Components | Angiomotin,E3 ubiquitin-protein ligase NEDD4-like | ||||||
 Keywords Keywords |  LIGASE / LIGASE /  WW domain / PPxY binding / WW domain / PPxY binding /  E3 Ubiquitin ligase / E3 Ubiquitin ligase /  Nedd4L Nedd4L | ||||||
| Function / homology |  Function and homology information Function and homology informationestablishment of cell polarity involved in ameboidal cell migration / positive regulation of caveolin-mediated endocytosis / RING-type E3 ubiquitin transferase (cysteine targeting) / negative regulation of sodium ion transmembrane transport / negative regulation of potassium ion transmembrane transporter activity / cell migration involved in gastrulation / regulation of potassium ion transmembrane transporter activity / negative regulation of potassium ion transmembrane transport / positive regulation of embryonic development / Regulation of CDH11 function ...establishment of cell polarity involved in ameboidal cell migration / positive regulation of caveolin-mediated endocytosis / RING-type E3 ubiquitin transferase (cysteine targeting) / negative regulation of sodium ion transmembrane transport / negative regulation of potassium ion transmembrane transporter activity / cell migration involved in gastrulation / regulation of potassium ion transmembrane transporter activity / negative regulation of potassium ion transmembrane transport / positive regulation of embryonic development / Regulation of CDH11 function / regulation of sodium ion transmembrane transport / blood vessel endothelial cell migration / negative regulation of sodium ion transmembrane transporter activity / establishment of epithelial cell polarity / negative regulation of protein localization to cell surface / regulation of membrane repolarization /  angiostatin binding / hippo signaling / positive regulation of dendrite extension / potassium channel inhibitor activity / ventricular cardiac muscle cell action potential / HECT-type E3 ubiquitin transferase / gastrulation with mouth forming second / negative regulation of vascular permeability / angiostatin binding / hippo signaling / positive regulation of dendrite extension / potassium channel inhibitor activity / ventricular cardiac muscle cell action potential / HECT-type E3 ubiquitin transferase / gastrulation with mouth forming second / negative regulation of vascular permeability /  sodium channel inhibitor activity / Signaling by Hippo / regulation of monoatomic ion transmembrane transport / cell-cell junction assembly / regulation of small GTPase mediated signal transduction / regulation of dendrite morphogenesis / sodium channel inhibitor activity / Signaling by Hippo / regulation of monoatomic ion transmembrane transport / cell-cell junction assembly / regulation of small GTPase mediated signal transduction / regulation of dendrite morphogenesis /  regulation of membrane depolarization / protein monoubiquitination / positive regulation of cell size / endocytic vesicle / sodium channel regulator activity / potassium channel regulator activity / bicellular tight junction / positive regulation of blood vessel endothelial cell migration / regulation of membrane depolarization / protein monoubiquitination / positive regulation of cell size / endocytic vesicle / sodium channel regulator activity / potassium channel regulator activity / bicellular tight junction / positive regulation of blood vessel endothelial cell migration /  vasculogenesis / protein K48-linked ubiquitination / vasculogenesis / protein K48-linked ubiquitination /  stress fiber / stress fiber /  regulation of cell migration / positive regulation of stress fiber assembly / monoatomic ion transmembrane transport / ruffle / regulation of cell migration / positive regulation of stress fiber assembly / monoatomic ion transmembrane transport / ruffle /  multivesicular body / negative regulation of angiogenesis / Downregulation of TGF-beta receptor signaling / multivesicular body / negative regulation of angiogenesis / Downregulation of TGF-beta receptor signaling /  regulation of membrane potential / regulation of membrane potential /  actin filament / Downregulation of SMAD2/3:SMAD4 transcriptional activity / Budding and maturation of HIV virion / actin filament / Downregulation of SMAD2/3:SMAD4 transcriptional activity / Budding and maturation of HIV virion /  regulation of protein stability / regulation of protein stability /  protein localization / Stimuli-sensing channels / positive regulation of protein catabolic process / ubiquitin-protein transferase activity / protein localization / Stimuli-sensing channels / positive regulation of protein catabolic process / ubiquitin-protein transferase activity /  chemotaxis / chemotaxis /  ubiquitin protein ligase activity / Antigen processing: Ubiquitination & Proteasome degradation / ubiquitin protein ligase activity / Antigen processing: Ubiquitination & Proteasome degradation /  cell junction / cell junction /  lamellipodium / lamellipodium /  signaling receptor activity / cytoplasmic vesicle / actin cytoskeleton organization / ubiquitin-dependent protein catabolic process / proteasome-mediated ubiquitin-dependent protein catabolic process / signaling receptor activity / cytoplasmic vesicle / actin cytoskeleton organization / ubiquitin-dependent protein catabolic process / proteasome-mediated ubiquitin-dependent protein catabolic process /  angiogenesis / in utero embryonic development / transmembrane transporter binding / angiogenesis / in utero embryonic development / transmembrane transporter binding /  cell differentiation / protein ubiquitination / external side of plasma membrane / cell differentiation / protein ubiquitination / external side of plasma membrane /  Golgi apparatus / Golgi apparatus /  cell surface / extracellular exosome / cell surface / extracellular exosome /  nucleoplasm / nucleoplasm /  membrane / membrane /  plasma membrane / plasma membrane /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / DGSA-distance geometry simulated annealing SOLUTION NMR / DGSA-distance geometry simulated annealing | ||||||
 Authors Authors | Alam, S.L. / Alian, A. / Thompson, T. / Rheinemann, L. / Volkman, B.F. / Peterson, F.C. / Sundquist, W.I. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2021 Journal: J.Biol.Chem. / Year: 2021Title: Interactions between AMOT PPxY motifs and NEDD4L WW domains function in HIV-1 release. Authors: Rheinemann, L. / Thompson, T. / Mercenne, G. / Paine, E.L. / Peterson, F.C. / Volkman, B.F. / Alam, S.L. / Alian, A. / Sundquist, W.I. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7lp5.cif.gz 7lp5.cif.gz | 451.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7lp5.ent.gz pdb7lp5.ent.gz | 380.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7lp5.json.gz 7lp5.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/lp/7lp5 https://data.pdbj.org/pub/pdb/validation_reports/lp/7lp5 ftp://data.pdbj.org/pub/pdb/validation_reports/lp/7lp5 ftp://data.pdbj.org/pub/pdb/validation_reports/lp/7lp5 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  7lp1C  7lp2C  7lp3C  7lp4C C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 7756.744 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: AMOT, KIAA1071, NEDD4L, KIAA0439, NEDL3 Homo sapiens (human) / Gene: AMOT, KIAA1071, NEDD4L, KIAA0439, NEDL3Production host:   Escherichia coli 'BL21-Gold(DE3)pLysS AG' (bacteria) Escherichia coli 'BL21-Gold(DE3)pLysS AG' (bacteria)References: UniProt: Q4VCS5, UniProt: Q96PU5, HECT-type E3 ubiquitin transferase |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||||||
| Sample conditions | Ionic strength: 50 mM / Label: 13C/15N_sample / pH: 7.5 / Pressure: 1 atm / Temperature: 298 K |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: DGSA-distance geometry simulated annealing / Software ordinal: 1 | ||||||||||||||||||||||||||||
| NMR representative | Selection criteria: target function | ||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: target function / Conformers calculated total number: 100 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller








 PDBj
PDBj







