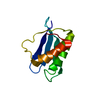
Entry Database : PDB / ID : 7cqpTitle Complex of TRPC4 and Calmodulin_Nlobe Calmodulin-1 Peptide from Short transient receptor potential channel 4 Keywords / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Mus musculus (house mouse)Method / / / / Resolution : 1.9 Å Authors Shen, Z.S. Funding support Organization Grant number Country National Natural Science Foundation of China (NSFC) 31670765 National Natural Science Foundation of China (NSFC) 31870746
Journal : Structure / Year : 2021Title : Calmodulin binds to Drosophila TRP with an unexpected mode.Authors : Chen, W. / Shen, Z. / Asteriti, S. / Chen, Z. / Ye, F. / Sun, Z. / Wan, J. / Montell, C. / Hardie, R.C. / Liu, W. / Zhang, M. History Deposition Aug 11, 2020 Deposition site / Processing site Revision 1.0 Jun 23, 2021 Provider / Type Revision 1.1 Nov 29, 2023 Group / Database references / Refinement descriptionCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model Item / _database_2.pdbx_database_accession
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords SIGNALING PROTEIN /
SIGNALING PROTEIN /  Calmodulin /
Calmodulin /  TRP channel
TRP channel Function and homology information
Function and homology information Protein methylation / RAF activation / VEGFR2 mediated vascular permeability / RAS processing / Smooth Muscle Contraction / gamma-aminobutyric acid secretion / Ca2+ pathway / FCERI mediated Ca+2 mobilization / RAF/MAP kinase cascade / RHO GTPases activate IQGAPs / Extra-nuclear estrogen signaling /
Protein methylation / RAF activation / VEGFR2 mediated vascular permeability / RAS processing / Smooth Muscle Contraction / gamma-aminobutyric acid secretion / Ca2+ pathway / FCERI mediated Ca+2 mobilization / RAF/MAP kinase cascade / RHO GTPases activate IQGAPs / Extra-nuclear estrogen signaling /  store-operated calcium channel activity / PKA activation / Platelet degranulation /
store-operated calcium channel activity / PKA activation / Platelet degranulation /  cation channel complex /
cation channel complex /  TRP channels / Stimuli-sensing channels / Ion homeostasis / type 3 metabotropic glutamate receptor binding /
TRP channels / Stimuli-sensing channels / Ion homeostasis / type 3 metabotropic glutamate receptor binding /  inositol 1,4,5 trisphosphate binding / calcium ion import / organelle localization by membrane tethering / mitochondrion-endoplasmic reticulum membrane tethering / regulation of cardiac muscle cell action potential / autophagosome membrane docking / cortical cytoskeleton /
inositol 1,4,5 trisphosphate binding / calcium ion import / organelle localization by membrane tethering / mitochondrion-endoplasmic reticulum membrane tethering / regulation of cardiac muscle cell action potential / autophagosome membrane docking / cortical cytoskeleton /  nitric-oxide synthase binding / oligodendrocyte differentiation / protein phosphatase activator activity / regulation of calcium ion transport /
nitric-oxide synthase binding / oligodendrocyte differentiation / protein phosphatase activator activity / regulation of calcium ion transport /  adenylate cyclase binding /
adenylate cyclase binding /  catalytic complex / detection of calcium ion / negative regulation of ryanodine-sensitive calcium-release channel activity / regulation of cardiac muscle contraction / calcium channel inhibitor activity / cellular response to interferon-beta /
catalytic complex / detection of calcium ion / negative regulation of ryanodine-sensitive calcium-release channel activity / regulation of cardiac muscle contraction / calcium channel inhibitor activity / cellular response to interferon-beta /  phosphatidylinositol 3-kinase binding / regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / positive regulation of protein dephosphorylation /
phosphatidylinositol 3-kinase binding / regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / positive regulation of protein dephosphorylation /  voltage-gated potassium channel complex / regulation of ryanodine-sensitive calcium-release channel activity /
voltage-gated potassium channel complex / regulation of ryanodine-sensitive calcium-release channel activity /  titin binding / sperm midpiece /
titin binding / sperm midpiece /  calcium channel complex / regulation of cytosolic calcium ion concentration / adenylate cyclase activator activity / nitric-oxide synthase regulator activity /
calcium channel complex / regulation of cytosolic calcium ion concentration / adenylate cyclase activator activity / nitric-oxide synthase regulator activity /  regulation of heart rate /
regulation of heart rate /  sarcomere / protein serine/threonine kinase activator activity /
sarcomere / protein serine/threonine kinase activator activity /  regulation of cytokinesis /
regulation of cytokinesis /  caveola / calcium ion transmembrane transport / positive regulation of receptor signaling pathway via JAK-STAT /
caveola / calcium ion transmembrane transport / positive regulation of receptor signaling pathway via JAK-STAT /  calcium channel activity / spindle microtubule /
calcium channel activity / spindle microtubule /  beta-catenin binding / cellular response to type II interferon /
beta-catenin binding / cellular response to type II interferon /  spindle pole / calcium ion transport / calcium-dependent protein binding / G2/M transition of mitotic cell cycle / cell-cell junction /
spindle pole / calcium ion transport / calcium-dependent protein binding / G2/M transition of mitotic cell cycle / cell-cell junction /  myelin sheath /
myelin sheath /  growth cone / basolateral plasma membrane / transmembrane transporter binding /
growth cone / basolateral plasma membrane / transmembrane transporter binding /  cadherin binding /
cadherin binding /  membrane raft / protein domain specific binding /
membrane raft / protein domain specific binding /  centrosome /
centrosome /  calcium ion binding /
calcium ion binding /  protein kinase binding /
protein kinase binding /  cell surface / protein-containing complex /
cell surface / protein-containing complex /  nucleoplasm /
nucleoplasm /  plasma membrane /
plasma membrane /  cytosol /
cytosol /  cytoplasm
cytoplasm
 Mus musculus (house mouse)
Mus musculus (house mouse) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT /
MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 1.9 Å
molecular replacement / Resolution: 1.9 Å  Authors
Authors China, 2items
China, 2items  Citation
Citation Journal: Structure / Year: 2021
Journal: Structure / Year: 2021 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 7cqp.cif.gz
7cqp.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb7cqp.ent.gz
pdb7cqp.ent.gz PDB format
PDB format 7cqp.json.gz
7cqp.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/cq/7cqp
https://data.pdbj.org/pub/pdb/validation_reports/cq/7cqp ftp://data.pdbj.org/pub/pdb/validation_reports/cq/7cqp
ftp://data.pdbj.org/pub/pdb/validation_reports/cq/7cqp


 Links
Links Assembly
Assembly
 Components
Components

 Mus musculus (house mouse) / Gene: Calm1, Calm, Cam, Cam1 / Production host:
Mus musculus (house mouse) / Gene: Calm1, Calm, Cam, Cam1 / Production host: 
 Escherichia coli (E. coli) / References: UniProt: P0DP26
Escherichia coli (E. coli) / References: UniProt: P0DP26 / TrpC4 / Capacitative calcium entry channel Trp4 / Receptor-activated cation channel TRP4
/ TrpC4 / Capacitative calcium entry channel Trp4 / Receptor-activated cation channel TRP4
 Mus musculus (house mouse) / References: UniProt: Q9QUQ5
Mus musculus (house mouse) / References: UniProt: Q9QUQ5 Water
Water X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation
 SYNCHROTRON / Site:
SYNCHROTRON / Site:  SSRF
SSRF  / Beamline: BL19U1 / Wavelength: 0.9789 Å
/ Beamline: BL19U1 / Wavelength: 0.9789 Å : 0.9789 Å / Relative weight: 1
: 0.9789 Å / Relative weight: 1 
 molecular replacement
molecular replacement Processing
Processing :
:  MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller












 PDBj
PDBj















