[English] 日本語
 Yorodumi
Yorodumi- PDB-6xs5: Crystal structure of human Vps29 complexed with RaPID-derived cyc... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6xs5 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of human Vps29 complexed with RaPID-derived cyclic peptide RT-D1 | ||||||
 Components Components |
| ||||||
 Keywords Keywords | PROTEIN TRANSPORT/INHIBITOR /  Vps29 / Vps29 /  Retromer / Retromer /  Endosome / Endosome /  Protein transport / Protein transport /  cyclic peptide / cyclic peptide /  inhibitor / PROTEIN TRANSPORT-INHIBITOR complex inhibitor / PROTEIN TRANSPORT-INHIBITOR complex | ||||||
| Function / homology |  Function and homology information Function and homology information retromer, cargo-selective complex / WNT ligand biogenesis and trafficking / retromer, cargo-selective complex / WNT ligand biogenesis and trafficking /  retromer complex / endocytic recycling / retromer complex / endocytic recycling /  retrograde transport, endosome to Golgi / retrograde transport, endosome to Golgi /  intracellular protein transport / late endosome / intracellular protein transport / late endosome /  early endosome / endosome membrane / early endosome / endosome membrane /  endosome ... endosome ... retromer, cargo-selective complex / WNT ligand biogenesis and trafficking / retromer, cargo-selective complex / WNT ligand biogenesis and trafficking /  retromer complex / endocytic recycling / retromer complex / endocytic recycling /  retrograde transport, endosome to Golgi / retrograde transport, endosome to Golgi /  intracellular protein transport / late endosome / intracellular protein transport / late endosome /  early endosome / endosome membrane / early endosome / endosome membrane /  endosome / intracellular membrane-bounded organelle / endosome / intracellular membrane-bounded organelle /  metal ion binding / metal ion binding /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human)synthetic construct (others) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 2.01 Å molecular replacement / Resolution: 2.01 Å | ||||||
 Authors Authors | Chen, K.-E. / Guo, Q. / Collins, B.M. | ||||||
| Funding support |  Australia, 1items Australia, 1items
| ||||||
 Citation Citation |  Journal: Sci Adv / Year: 2021 Journal: Sci Adv / Year: 2021Title: De novo macrocyclic peptides for inhibiting, stabilizing, and probing the function of the retromer endosomal trafficking complex. Authors: Kai-En Chen / Qian Guo / Timothy A Hill / Yi Cui / Amy K Kendall / Zhe Yang / Ryan J Hall / Michael D Healy / Joanna Sacharz / Suzanne J Norwood / Sachini Fonseka / Boyang Xie / Robert C ...Authors: Kai-En Chen / Qian Guo / Timothy A Hill / Yi Cui / Amy K Kendall / Zhe Yang / Ryan J Hall / Michael D Healy / Joanna Sacharz / Suzanne J Norwood / Sachini Fonseka / Boyang Xie / Robert C Reid / Natalya Leneva / Robert G Parton / Rajesh Ghai / David A Stroud / David P Fairlie / Hiroaki Suga / Lauren P Jackson / Rohan D Teasdale / Toby Passioura / Brett M Collins /    Abstract: The retromer complex (Vps35-Vps26-Vps29) is essential for endosomal membrane trafficking and signaling. Mutation of the retromer subunit Vps35 causes late-onset Parkinson’s disease, while viral and ...The retromer complex (Vps35-Vps26-Vps29) is essential for endosomal membrane trafficking and signaling. Mutation of the retromer subunit Vps35 causes late-onset Parkinson’s disease, while viral and bacterial pathogens can hijack the complex during cellular infection. To modulate and probe its function, we have created a novel series of macrocyclic peptides that bind retromer with high affinity and specificity. Crystal structures show that most of the cyclic peptides bind to Vps29 via a Pro-Leu–containing sequence, structurally mimicking known interactors such as TBC1D5 and blocking their interaction with retromer in vitro and in cells. By contrast, macrocyclic peptide RT-L4 binds retromer at the Vps35-Vps26 interface and is a more effective molecular chaperone than reported small molecules, suggesting a new therapeutic avenue for targeting retromer. Last, tagged peptides can be used to probe the cellular localization of retromer and its functional interactions in cells, providing novel tools for studying retromer function. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6xs5.cif.gz 6xs5.cif.gz | 61.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6xs5.ent.gz pdb6xs5.ent.gz | 40.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6xs5.json.gz 6xs5.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/xs/6xs5 https://data.pdbj.org/pub/pdb/validation_reports/xs/6xs5 ftp://data.pdbj.org/pub/pdb/validation_reports/xs/6xs5 ftp://data.pdbj.org/pub/pdb/validation_reports/xs/6xs5 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 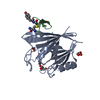 6xs7C  6xs8C  6xs9C  6xsaC  1w24S S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||
| Unit cell |
|
- Components
Components
| #1: Protein |  Vacuole / hVPS29 / PEP11 homolog / Vesicle protein sorting 29 Vacuole / hVPS29 / PEP11 homolog / Vesicle protein sorting 29Mass: 21636.869 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: VPS29, DC15, DC7, MDS007 / Plasmid: pGEX-4T2 / Production host: Homo sapiens (human) / Gene: VPS29, DC15, DC7, MDS007 / Plasmid: pGEX-4T2 / Production host:   Escherichia coli BL21(DE3) (bacteria) / References: UniProt: Q9UBQ0 Escherichia coli BL21(DE3) (bacteria) / References: UniProt: Q9UBQ0 | ||||||
|---|---|---|---|---|---|---|---|
| #2: Protein/peptide | Mass: 1984.364 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: Cyclic peptide RT-D1 derived from RaPID screen / Source: (synth.) synthetic construct (others) | ||||||
| #3: Chemical |  Glycerol Glycerol#4: Chemical | ChemComp-FMT / |  Formic acid Formic acid#5: Water | ChemComp-HOH / |  Water WaterHas ligand of interest | Y | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.25 Å3/Da / Density % sol: 45.27 % |
|---|---|
Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / Details: 3.5 M Sodium Formate |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Australian Synchrotron Australian Synchrotron  / Beamline: MX2 / Wavelength: 0.953729 Å / Beamline: MX2 / Wavelength: 0.953729 Å |
| Detector | Type: DECTRIS EIGER X 16M / Detector: PIXEL / Date: Sep 23, 2018 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.953729 Å / Relative weight: 1 : 0.953729 Å / Relative weight: 1 |
| Reflection | Resolution: 2.01→42.41 Å / Num. obs: 14169 / % possible obs: 99.9 % / Redundancy: 6.9 % / Biso Wilson estimate: 26.373 Å2 / CC1/2: 0.999 / Rmerge(I) obs: 0.077 / Rpim(I) all: 0.032 / Rrim(I) all: 0.083 / Net I/σ(I): 12.2 |
| Reflection shell | Resolution: 2.01→2.06 Å / Redundancy: 7 % / Rmerge(I) obs: 0.585 / Mean I/σ(I) obs: 2.5 / Num. unique obs: 1030 / CC1/2: 0.917 / Rpim(I) all: 0.235 / Rrim(I) all: 0.632 / % possible all: 98.8 |
-Phasing
Phasing | Method:  molecular replacement molecular replacement | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Phasing MR | Model details: Phaser MODE: MR_AUTO
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1W24 Resolution: 2.01→42.41 Å / SU ML: 0.205837110947 / Cross valid method: FREE R-VALUE / σ(F): 1.37769675716 / Phase error: 25.274220866 Stereochemistry target values: GeoStd + Monomer Library + CDL v1.2
| ||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 32.3112114153 Å2 | ||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.01→42.41 Å
| ||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller














 PDBj
PDBj





