[English] 日本語
 Yorodumi
Yorodumi- PDB-6pti: STRUCTURE OF FORM III CRYSTALS OF BOVINE PANCREATIC TRYPSIN INHIBITOR -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6pti | ||||||
|---|---|---|---|---|---|---|---|
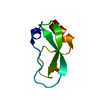
| Title | STRUCTURE OF FORM III CRYSTALS OF BOVINE PANCREATIC TRYPSIN INHIBITOR | ||||||
 Components Components | PANCREATIC TRYPSIN INHIBITOR PRECURSOR | ||||||
 Keywords Keywords | PROTEINASE INHIBITOR (TRYPSIN) | ||||||
| Function / homology |  Function and homology information Function and homology informationtrypsinogen activation / negative regulation of serine-type endopeptidase activity /  sulfate binding / potassium channel inhibitor activity / negative regulation of platelet aggregation / sulfate binding / potassium channel inhibitor activity / negative regulation of platelet aggregation /  zymogen binding / molecular function inhibitor activity / negative regulation of thrombin-activated receptor signaling pathway / zymogen binding / molecular function inhibitor activity / negative regulation of thrombin-activated receptor signaling pathway /  serine protease inhibitor complex / serine-type endopeptidase inhibitor activity ...trypsinogen activation / negative regulation of serine-type endopeptidase activity / serine protease inhibitor complex / serine-type endopeptidase inhibitor activity ...trypsinogen activation / negative regulation of serine-type endopeptidase activity /  sulfate binding / potassium channel inhibitor activity / negative regulation of platelet aggregation / sulfate binding / potassium channel inhibitor activity / negative regulation of platelet aggregation /  zymogen binding / molecular function inhibitor activity / negative regulation of thrombin-activated receptor signaling pathway / zymogen binding / molecular function inhibitor activity / negative regulation of thrombin-activated receptor signaling pathway /  serine protease inhibitor complex / serine-type endopeptidase inhibitor activity / serine protease inhibitor complex / serine-type endopeptidase inhibitor activity /  protease binding / protease binding /  calcium ion binding / calcium ion binding /  extracellular space extracellular spaceSimilarity search - Function | ||||||
| Biological species |   Bos taurus (cattle) Bos taurus (cattle) | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 1.7 Å X-RAY DIFFRACTION / Resolution: 1.7 Å | ||||||
 Authors Authors | Wlodawer, A. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1987 Journal: J.Mol.Biol. / Year: 1987Title: Structure of form III crystals of bovine pancreatic trypsin inhibitor. Authors: Wlodawer, A. / Nachman, J. / Gilliland, G.L. / Gallagher, W. / Woodward, C. #1:  Journal: J.Mol.Biol. / Year: 1984 Journal: J.Mol.Biol. / Year: 1984Title: Structure of Bovine Pancreatic Trypsin Inhibitor. Results of Joint Neutron and X-Ray Refinement of Crystal Form II Authors: Wlodawer, A. / Walter, J. / Huber, R. / Sjolin, L. #2:  Journal: J.Mol.Biol. / Year: 1987 Journal: J.Mol.Biol. / Year: 1987Title: Comparison of Two Highly Refined Structures of Bovine Pancreatic Trypsin Inhibitor Authors: Wlodawer, A. / Deisenhofer, J. / Huber, R. #3:  Journal: J.Mol.Biol. / Year: 1983 Journal: J.Mol.Biol. / Year: 1983Title: Pancreatic Trypsin Inhibitor. A New Crystal Form and its Analysis Authors: Walter, J. / Huber, R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6pti.cif.gz 6pti.cif.gz | 24.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6pti.ent.gz pdb6pti.ent.gz | 15.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6pti.json.gz 6pti.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pt/6pti https://data.pdbj.org/pub/pdb/validation_reports/pt/6pti ftp://data.pdbj.org/pub/pdb/validation_reports/pt/6pti ftp://data.pdbj.org/pub/pdb/validation_reports/pt/6pti | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||
| Unit cell |
| ||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 6527.568 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Bos taurus (cattle) / References: UniProt: P00974 Bos taurus (cattle) / References: UniProt: P00974 |
|---|---|
| #2: Chemical | ChemComp-PO4 /  Phosphate Phosphate |
| #3: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.94 Å3/Da / Density % sol: 36.63 % | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | *PLUS Temperature: 0 K / pH: 10.13 / Method: batch method | ||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Reflection | *PLUS Highest resolution: 1.7 Å / Num. all: 5753 / Num. obs: 5218 / Num. measured all: 22389 / Rmerge(I) obs: 0.04 |
|---|
- Processing
Processing
| Software | Name: PROLSQ / Classification: refinement | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Rfactor obs: 0.16 / Highest resolution: 1.7 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 1.7 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.16 / Highest resolution: 1.7 Å / Num. reflection obs: 4980 / σ(F): 1.5 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller











 PDBj
PDBj






