+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6jjx | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal Structure of KIBRA and Angiomotin complex | ||||||
 Components Components |
| ||||||
 Keywords Keywords |  SIGNALING PROTEIN / WW tandem / PY tandem / SIGNALING PROTEIN / WW tandem / PY tandem /  KIBRA / KIBRA /  Angiomotin / AMOT Angiomotin / AMOT | ||||||
| Function / homology |  Function and homology information Function and homology informationestablishment of cell polarity involved in ameboidal cell migration / regulation of hippo signaling / Signaling by Hippo / cell migration involved in gastrulation / positive regulation of embryonic development / Regulation of CDH11 function / blood vessel endothelial cell migration / establishment of epithelial cell polarity / negative regulation of organ growth /  angiostatin binding ...establishment of cell polarity involved in ameboidal cell migration / regulation of hippo signaling / Signaling by Hippo / cell migration involved in gastrulation / positive regulation of embryonic development / Regulation of CDH11 function / blood vessel endothelial cell migration / establishment of epithelial cell polarity / negative regulation of organ growth / angiostatin binding ...establishment of cell polarity involved in ameboidal cell migration / regulation of hippo signaling / Signaling by Hippo / cell migration involved in gastrulation / positive regulation of embryonic development / Regulation of CDH11 function / blood vessel endothelial cell migration / establishment of epithelial cell polarity / negative regulation of organ growth /  angiostatin binding / hippo signaling / gastrulation with mouth forming second / negative regulation of vascular permeability / Signaling by Hippo / cell-cell junction assembly / regulation of small GTPase mediated signal transduction / negative regulation of hippo signaling / positive regulation of cell size / endocytic vesicle / bicellular tight junction / positive regulation of blood vessel endothelial cell migration / angiostatin binding / hippo signaling / gastrulation with mouth forming second / negative regulation of vascular permeability / Signaling by Hippo / cell-cell junction assembly / regulation of small GTPase mediated signal transduction / negative regulation of hippo signaling / positive regulation of cell size / endocytic vesicle / bicellular tight junction / positive regulation of blood vessel endothelial cell migration /  vasculogenesis / vasculogenesis /  stress fiber / stress fiber /  regulation of cell migration / positive regulation of stress fiber assembly / ruffle / negative regulation of angiogenesis / regulation of cell migration / positive regulation of stress fiber assembly / ruffle / negative regulation of angiogenesis /  actin filament / actin filament /  protein localization / protein localization /  kinase binding / ruffle membrane / kinase binding / ruffle membrane /  chemotaxis / chemotaxis /  cell migration / cell migration /  cell junction / cell junction /  lamellipodium / protein-macromolecule adaptor activity / lamellipodium / protein-macromolecule adaptor activity /  signaling receptor activity / cytoplasmic vesicle / actin cytoskeleton organization / signaling receptor activity / cytoplasmic vesicle / actin cytoskeleton organization /  angiogenesis / in utero embryonic development / positive regulation of MAPK cascade / angiogenesis / in utero embryonic development / positive regulation of MAPK cascade /  transcription coactivator activity / molecular adaptor activity / negative regulation of cell population proliferation / external side of plasma membrane / regulation of DNA-templated transcription / perinuclear region of cytoplasm / negative regulation of transcription by RNA polymerase II / transcription coactivator activity / molecular adaptor activity / negative regulation of cell population proliferation / external side of plasma membrane / regulation of DNA-templated transcription / perinuclear region of cytoplasm / negative regulation of transcription by RNA polymerase II /  cell surface / protein-containing complex / cell surface / protein-containing complex /  nucleoplasm / nucleoplasm /  membrane / membrane /  nucleus / nucleus /  plasma membrane / plasma membrane /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Mus musculus (house mouse) Mus musculus (house mouse)  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | ||||||
 Authors Authors | Lin, Z. / Yang, Z. / Ji, Z. / Zhang, M. | ||||||
 Citation Citation |  Journal: Elife / Year: 2019 Journal: Elife / Year: 2019Title: Decoding WW domain tandem-mediated target recognitions in tissue growth and cell polarity. Authors: Lin, Z. / Yang, Z. / Xie, R. / Ji, Z. / Guan, K. / Zhang, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6jjx.cif.gz 6jjx.cif.gz | 135.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6jjx.ent.gz pdb6jjx.ent.gz | 105.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6jjx.json.gz 6jjx.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/jj/6jjx https://data.pdbj.org/pub/pdb/validation_reports/jj/6jjx ftp://data.pdbj.org/pub/pdb/validation_reports/jj/6jjx ftp://data.pdbj.org/pub/pdb/validation_reports/jj/6jjx | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6j68SC 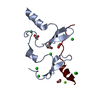 6jjwC  6jjyC  6jjzC  6jk0C  6jk1C S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 16036.611 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Mus musculus (house mouse) / Gene: Wwc1, Kiaa0869 Mus musculus (house mouse) / Gene: Wwc1, Kiaa0869Production host:   Escherichia coli str. 'clone D i2' (bacteria) Escherichia coli str. 'clone D i2' (bacteria)References: UniProt: Q5SXA9 #2: Protein/peptide |  Mass: 2788.063 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: AMOT, KIAA1071 / Production host: Homo sapiens (human) / Gene: AMOT, KIAA1071 / Production host:   Escherichia coli (E. coli) / References: UniProt: Q4VCS5 Escherichia coli (E. coli) / References: UniProt: Q4VCS5#3: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.5 Å3/Da / Density % sol: 50.8 % |
|---|---|
Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop Details: 25% w/v pentaerythritol propoxylat 629 (17/8 PO/OH), 50mM MgCl2, 100mM Tris (pH 8.5) PH range: 8.5 |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRF SSRF  / Beamline: BL19U1 / Wavelength: 0.97853 Å / Beamline: BL19U1 / Wavelength: 0.97853 Å |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Mar 18, 2016 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.97853 Å / Relative weight: 1 : 0.97853 Å / Relative weight: 1 |
| Reflection | Resolution: 2→50 Å / Num. obs: 26165 / % possible obs: 99.8 % / Redundancy: 9.9 % / Rmerge(I) obs: 0.103 / Net I/σ(I): 22.4 |
| Reflection shell | Resolution: 2→2.03 Å / Redundancy: 10.2 % / Rmerge(I) obs: 0.752 / Mean I/σ(I) obs: 2.2 / Num. unique obs: 1282 / % possible all: 99.9 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 6J68 Resolution: 2→39.687 Å / SU ML: 0.21 / Cross valid method: FREE R-VALUE / σ(F): 1.38 / Phase error: 26.12
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→39.687 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 65.3741 Å / Origin y: 27.5841 Å / Origin z: 99.9216 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: all |
 Movie
Movie Controller
Controller









 PDBj
PDBj





