[English] 日本語
 Yorodumi
Yorodumi- PDB-5n7e: Crystal structure of the Dbl-homology domain of Bcr-Abl in comple... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5n7e | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of the Dbl-homology domain of Bcr-Abl in complex with monobody Mb(Bcr-DH_4). | ||||||
 Components Components |
| ||||||
 Keywords Keywords |  SIGNALING PROTEIN / Dbl-homology / SIGNALING PROTEIN / Dbl-homology /  monobody / monobody /  Bcr-Abl / Bcr-Abl /  transferase transferase | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of respiratory burst / negative regulation of cellular extravasation / negative regulation of macrophage migration / : / negative regulation of blood vessel remodeling / negative regulation of neutrophil degranulation / macrophage migration / neutrophil degranulation /  intracellular protein transmembrane transport / intracellular protein transmembrane transport /  renal system process ...negative regulation of respiratory burst / negative regulation of cellular extravasation / negative regulation of macrophage migration / : / negative regulation of blood vessel remodeling / negative regulation of neutrophil degranulation / macrophage migration / neutrophil degranulation / renal system process ...negative regulation of respiratory burst / negative regulation of cellular extravasation / negative regulation of macrophage migration / : / negative regulation of blood vessel remodeling / negative regulation of neutrophil degranulation / macrophage migration / neutrophil degranulation /  intracellular protein transmembrane transport / intracellular protein transmembrane transport /  renal system process / renal system process /  regulation of vascular permeability / regulation of Rho protein signal transduction / regulation of vascular permeability / regulation of Rho protein signal transduction /  focal adhesion assembly / definitive hemopoiesis / Signaling by cytosolic FGFR1 fusion mutants / activation of GTPase activity / regulation of small GTPase mediated signal transduction / inner ear morphogenesis / small GTPase-mediated signal transduction / RHOB GTPase cycle / RHOC GTPase cycle / CDC42 GTPase cycle / neuromuscular process controlling balance / homeostasis of number of cells / RHOA GTPase cycle / negative regulation of reactive oxygen species metabolic process / RAC2 GTPase cycle / RAC3 GTPase cycle / focal adhesion assembly / definitive hemopoiesis / Signaling by cytosolic FGFR1 fusion mutants / activation of GTPase activity / regulation of small GTPase mediated signal transduction / inner ear morphogenesis / small GTPase-mediated signal transduction / RHOB GTPase cycle / RHOC GTPase cycle / CDC42 GTPase cycle / neuromuscular process controlling balance / homeostasis of number of cells / RHOA GTPase cycle / negative regulation of reactive oxygen species metabolic process / RAC2 GTPase cycle / RAC3 GTPase cycle /  phagocytosis / positive regulation of phagocytosis / keratinocyte differentiation / RAC1 GTPase cycle / Signaling by FGFR1 in disease / phagocytosis / positive regulation of phagocytosis / keratinocyte differentiation / RAC1 GTPase cycle / Signaling by FGFR1 in disease /  GTPase activator activity / guanyl-nucleotide exchange factor activity / Schaffer collateral - CA1 synapse / GTPase activator activity / guanyl-nucleotide exchange factor activity / Schaffer collateral - CA1 synapse /  brain development / modulation of chemical synaptic transmission / negative regulation of inflammatory response / actin cytoskeleton organization / brain development / modulation of chemical synaptic transmission / negative regulation of inflammatory response / actin cytoskeleton organization /  protein tyrosine kinase activity / cellular response to lipopolysaccharide / protein tyrosine kinase activity / cellular response to lipopolysaccharide /  dendritic spine / dendritic spine /  postsynaptic density / postsynaptic density /  non-specific serine/threonine protein kinase / non-specific serine/threonine protein kinase /  regulation of cell cycle / regulation of cell cycle /  axon / axon /  protein phosphorylation / protein serine kinase activity / protein serine/threonine kinase activity / glutamatergic synapse / protein phosphorylation / protein serine kinase activity / protein serine/threonine kinase activity / glutamatergic synapse /  signal transduction / protein-containing complex / extracellular exosome / signal transduction / protein-containing complex / extracellular exosome /  ATP binding / ATP binding /  membrane / membrane /  plasma membrane / plasma membrane /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species | synthetic construct (others)  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.647 Å MOLECULAR REPLACEMENT / Resolution: 1.647 Å | ||||||
 Authors Authors | Reckel, S. / Reynaud, A. / Pojer, F. / Hantschel, O. | ||||||
| Funding support |  Switzerland, 1items Switzerland, 1items
| ||||||
 Citation Citation |  Journal: Nat Commun / Year: 2017 Journal: Nat Commun / Year: 2017Title: Structural and functional dissection of the DH and PH domains of oncogenic Bcr-Abl tyrosine kinase. Authors: Sina Reckel / Charlotte Gehin / Delphine Tardivon / Sandrine Georgeon / Tim Kükenshöner / Frank Löhr / Akiko Koide / Lena Buchner / Alejandro Panjkovich / Aline Reynaud / Sara Pinho / ...Authors: Sina Reckel / Charlotte Gehin / Delphine Tardivon / Sandrine Georgeon / Tim Kükenshöner / Frank Löhr / Akiko Koide / Lena Buchner / Alejandro Panjkovich / Aline Reynaud / Sara Pinho / Barbara Gerig / Dmitri Svergun / Florence Pojer / Peter Güntert / Volker Dötsch / Shohei Koide / Anne-Claude Gavin / Oliver Hantschel /     Abstract: The two isoforms of the Bcr-Abl tyrosine kinase, p210 and p190, are associated with different leukemias and have a dramatically different signaling network, despite similar kinase activity. To ...The two isoforms of the Bcr-Abl tyrosine kinase, p210 and p190, are associated with different leukemias and have a dramatically different signaling network, despite similar kinase activity. To provide a molecular rationale for these observations, we study the Dbl-homology (DH) and Pleckstrin-homology (PH) domains of Bcr-Abl p210, which constitute the only structural differences to p190. Here we report high-resolution structures of the DH and PH domains and characterize conformations of the DH-PH unit in solution. Our structural and functional analyses show no evidence that the DH domain acts as a guanine nucleotide exchange factor, whereas the PH domain binds to various phosphatidylinositol-phosphates. PH-domain mutants alter subcellular localization and result in decreased interactions with p210-selective interaction partners. Hence, the PH domain, but not the DH domain, plays an important role in the formation of the differential p210 and p190 Bcr-Abl signaling networks. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5n7e.cif.gz 5n7e.cif.gz | 80.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5n7e.ent.gz pdb5n7e.ent.gz | 58.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5n7e.json.gz 5n7e.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/n7/5n7e https://data.pdbj.org/pub/pdb/validation_reports/n7/5n7e ftp://data.pdbj.org/pub/pdb/validation_reports/n7/5n7e ftp://data.pdbj.org/pub/pdb/validation_reports/n7/5n7e | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5n6rC  5oc7C  2z0qS 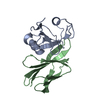 4je4S S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 10121.201 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.) synthetic construct (others) / Production host:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
|---|---|
| #2: Protein | Mass: 24901.482 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: BCR, BCR1, D22S11 / Plasmid: pETM-11 / Production host: Homo sapiens (human) / Gene: BCR, BCR1, D22S11 / Plasmid: pETM-11 / Production host:   Escherichia coli (E. coli) Escherichia coli (E. coli)References: UniProt: P11274,  non-specific serine/threonine protein kinase non-specific serine/threonine protein kinase |
| #3: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.83 Å3/Da / Density % sol: 56.47 % |
|---|---|
Crystal grow | Temperature: 291 K / Method: vapor diffusion, sitting drop / pH: 7.5 Details: 0.12 M monosaccharides (0.2M D-Glucose; 0.2M D-Mannose; 0.2M D-Galactose; 0.2M L-Fucose; 0.2M D-Xylose; 0.2M N-Acetyl-D-Glucosamine); 0.1 M buffer system 2 pH 7.5 (Sodium HEPES; MOPS (acid)); ...Details: 0.12 M monosaccharides (0.2M D-Glucose; 0.2M D-Mannose; 0.2M D-Galactose; 0.2M L-Fucose; 0.2M D-Xylose; 0.2M N-Acetyl-D-Glucosamine); 0.1 M buffer system 2 pH 7.5 (Sodium HEPES; MOPS (acid)); 50% precipitant mix 1 (40% v/v PEG 500* MME; 20 % w/v PEG 20000) |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SLS SLS  / Beamline: X06DA / Wavelength: 0.99987 Å / Beamline: X06DA / Wavelength: 0.99987 Å |
| Detector | Type: DECTRIS PILATUS 2M-F / Detector: PIXEL / Date: Aug 21, 2015 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.99987 Å / Relative weight: 1 : 0.99987 Å / Relative weight: 1 |
| Reflection | Resolution: 1.647→43.402 Å / Num. obs: 47055 / % possible obs: 99.27 % / Redundancy: 3.8 % / CC1/2: 0.999 / Rmerge(I) obs: 0.03673 / Rpim(I) all: 0.02165 / Net I/σ(I): 18.29 |
| Reflection shell | Resolution: 1.647→1.706 Å / Rmerge(I) obs: 0.5916 / Mean I/σ(I) all: 1.94 / Num. unique all: 4391 / CC1/2: 0.946 / Rpim(I) all: 0.3474 / % possible all: 93.25 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: Chain A: homology model based on 4JE4, Chain B: homology model based on 2Z0Q Resolution: 1.647→43.402 Å / SU ML: 0.21 / Cross valid method: NONE / σ(F): 1.35 / Phase error: 23.2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 101.18 Å2 / Biso mean: 36.1086 Å2 / Biso min: 15.49 Å2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 1.647→43.402 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Rfactor Rfree error: 0 / Total num. of bins used: 18
|
 Movie
Movie Controller
Controller















 PDBj
PDBj







