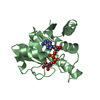
Entry Database : PDB / ID : 4z9aTitle Crystal structure of Low Molecular Weight Protein Tyrosine Phosphatase isoform A complexed with phenylmethanesulfonic acid Low molecular weight phosphotyrosine protein phosphatase Keywords / / / Function / homology Biological species Homo sapiens (human)Method / / / Resolution : 2.1 Å Authors Trivella, D.B.B. / Fonseca, E.M.B. / Scorsato, V. / Dias, M.P. / Aparicio, R. Funding support Organization Grant number Country Sao Paulo Research Foundation (FAPESP) 2009/51602-5 Sao Paulo Research Foundation (FAPESP) 2010/17544-5 Sao Paulo Research Foundation (FAPESP) 2011/15792-4 Sao Paulo Research Foundation (FAPESP) 2011/03054-9
Journal : Bioorg.Med.Chem. / Year : 2015Title : Crystal structures of the apo form and a complex of human LMW-PTP with a phosphonic acid provide new evidence of a secondary site potentially related to the anchorage of natural substrates.Authors : Fonseca, E.M. / Trivella, D.B. / Scorsato, V. / Dias, M.P. / Bazzo, N.L. / Mandapati, K.R. / de Oliveira, F.L. / Ferreira-Halder, C.V. / Pilli, R.A. / Miranda, P.C. / Aparicio, R. History Deposition Apr 10, 2015 Deposition site / Processing site Revision 1.0 Jul 15, 2015 Provider / Type Revision 1.1 Aug 5, 2015 Group Revision 1.2 Jan 17, 2018 Group / Database references / Derived calculationsCategory / pdbx_audit_support / pdbx_struct_oper_listItem / _pdbx_audit_support.funding_organization / _pdbx_struct_oper_list.symmetry_operationRevision 1.3 Apr 17, 2019 Group / Data collection / Category / Item Revision 1.4 Jan 1, 2020 Group / Category / Item Revision 1.5 Sep 27, 2023 Group / Database references / Refinement descriptionCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model Item / _database_2.pdbx_database_accession
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords HYDROLASE /
HYDROLASE /  Protein Tyrosine Phosphatase /
Protein Tyrosine Phosphatase /  protein-ligand complex / phenylmethanesulfonic acid
protein-ligand complex / phenylmethanesulfonic acid Function and homology information
Function and homology information acid phosphatase /
acid phosphatase /  acid phosphatase activity / non-membrane spanning protein tyrosine phosphatase activity /
acid phosphatase activity / non-membrane spanning protein tyrosine phosphatase activity /  protein-tyrosine-phosphatase /
protein-tyrosine-phosphatase /  protein tyrosine phosphatase activity /
protein tyrosine phosphatase activity /  sarcolemma / cytoplasmic side of plasma membrane / extracellular exosome /
sarcolemma / cytoplasmic side of plasma membrane / extracellular exosome /  cytosol /
cytosol /  cytoplasm
cytoplasm
 Homo sapiens (human)
Homo sapiens (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.1 Å
MOLECULAR REPLACEMENT / Resolution: 2.1 Å  Authors
Authors Brazil, 4items
Brazil, 4items  Citation
Citation Journal: Bioorg.Med.Chem. / Year: 2015
Journal: Bioorg.Med.Chem. / Year: 2015 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 4z9a.cif.gz
4z9a.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb4z9a.ent.gz
pdb4z9a.ent.gz PDB format
PDB format 4z9a.json.gz
4z9a.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/z9/4z9a
https://data.pdbj.org/pub/pdb/validation_reports/z9/4z9a ftp://data.pdbj.org/pub/pdb/validation_reports/z9/4z9a
ftp://data.pdbj.org/pub/pdb/validation_reports/z9/4z9a


 Links
Links Assembly
Assembly
 Components
Components
 Homo sapiens (human) / Gene: ACP1 / Plasmid: pET28a / Production host:
Homo sapiens (human) / Gene: ACP1 / Plasmid: pET28a / Production host: 
 Escherichia coli (E. coli) / Strain (production host): BL21DE3
Escherichia coli (E. coli) / Strain (production host): BL21DE3 protein-tyrosine-phosphatase,
protein-tyrosine-phosphatase,  acid phosphatase
acid phosphatase Glycerol
Glycerol Sulfate
Sulfate Water
Water X-RAY DIFFRACTION
X-RAY DIFFRACTION Sample preparation
Sample preparation
 SYNCHROTRON / Site:
SYNCHROTRON / Site:  LNLS
LNLS  / Beamline: W01B-MX2 / Wavelength: 1.459 Å
/ Beamline: W01B-MX2 / Wavelength: 1.459 Å : 1.459 Å / Relative weight: 1
: 1.459 Å / Relative weight: 1  Processing
Processing :
:  MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller












 PDBj
PDBj







