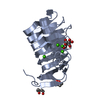
Entry Database : PDB / ID : 4yzxTitle C. bescii Family 3 pectate lyase double mutant K108A/D107N in complex with trigalacturonic acid Pectate lyase Keywords / / Function / homology Biological species Caldicellulosiruptor bescii (bacteria)Method / / Resolution : 1.25 Å Authors Alahuhta, P.M. / Lunin, V.V. Funding support Organization Grant number Country Department of Energy (DOE, United States)
Journal : Acta Crystallogr.,Sect.D / Year : 2015Title : The catalytic mechanism and unique low pH optimum of Caldicellulosiruptor bescii family 3 pectate lyase.Authors : Alahuhta, M. / Taylor, L.E. / Brunecky, R. / Sammond, D.W. / Michener, W. / Adams, M.W. / Himmel, M.E. / Bomble, Y.J. / Lunin, V. History Deposition Mar 25, 2015 Deposition site / Processing site Revision 1.0 Dec 23, 2015 Provider / Type Revision 1.1 Sep 20, 2017 Group / Derived calculations / Category / pdbx_struct_oper_listItem / _pdbx_struct_oper_list.symmetry_operationRevision 1.2 Dec 4, 2019 Group / Data collection / Category / pdbx_audit_supportItem / _pdbx_audit_support.funding_organizationRevision 2.0 Jul 29, 2020 Group Atomic model / Data collection ... Atomic model / Data collection / Derived calculations / Structure summary Category atom_site / atom_site_anisotrop ... atom_site / atom_site_anisotrop / chem_comp / entity / pdbx_branch_scheme / pdbx_chem_comp_identifier / pdbx_entity_branch / pdbx_entity_branch_descriptor / pdbx_entity_branch_link / pdbx_entity_branch_list / pdbx_entity_nonpoly / pdbx_nonpoly_scheme / pdbx_struct_assembly_gen / pdbx_struct_conn_angle / pdbx_struct_special_symmetry / struct_asym / struct_conn / struct_conn_type / struct_site / struct_site_gen Item _atom_site.B_iso_or_equiv / _atom_site.Cartn_x ... _atom_site.B_iso_or_equiv / _atom_site.Cartn_x / _atom_site.Cartn_y / _atom_site.Cartn_z / _atom_site.auth_asym_id / _atom_site.auth_atom_id / _atom_site.auth_comp_id / _atom_site.auth_seq_id / _atom_site.label_alt_id / _atom_site.label_asym_id / _atom_site.label_atom_id / _atom_site.label_comp_id / _atom_site.label_entity_id / _atom_site.occupancy / _atom_site.type_symbol / _atom_site_anisotrop.U[1][1] / _atom_site_anisotrop.U[1][2] / _atom_site_anisotrop.U[1][3] / _atom_site_anisotrop.U[2][2] / _atom_site_anisotrop.U[2][3] / _atom_site_anisotrop.U[3][3] / _atom_site_anisotrop.pdbx_auth_asym_id / _atom_site_anisotrop.pdbx_auth_atom_id / _atom_site_anisotrop.pdbx_auth_comp_id / _atom_site_anisotrop.pdbx_auth_seq_id / _atom_site_anisotrop.pdbx_label_alt_id / _atom_site_anisotrop.pdbx_label_asym_id / _atom_site_anisotrop.pdbx_label_atom_id / _atom_site_anisotrop.pdbx_label_comp_id / _atom_site_anisotrop.type_symbol / _chem_comp.name / _chem_comp.type / _pdbx_entity_nonpoly.comp_id / _pdbx_entity_nonpoly.entity_id / _pdbx_entity_nonpoly.name / _pdbx_struct_assembly_gen.asym_id_list / _pdbx_struct_conn_angle.ptnr1_auth_asym_id / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_alt_id / _pdbx_struct_conn_angle.ptnr1_label_asym_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr2_label_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_alt_id / _pdbx_struct_conn_angle.ptnr3_label_asym_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.value / _pdbx_struct_special_symmetry.label_asym_id / _struct_conn.conn_type_id / _struct_conn.id / _struct_conn.pdbx_dist_value / _struct_conn.pdbx_leaving_atom_flag / _struct_conn.pdbx_ptnr1_label_alt_id / _struct_conn.pdbx_ptnr2_label_alt_id / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn_type.id Description / Provider / Type Revision 2.1 Mar 6, 2024 Group Data collection / Database references ... Data collection / Database references / Derived calculations / Structure summary Category chem_comp / chem_comp_atom ... chem_comp / chem_comp_atom / chem_comp_bond / database_2 / struct_conn Item _chem_comp.pdbx_synonyms / _database_2.pdbx_DOI ... _chem_comp.pdbx_synonyms / _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _struct_conn.pdbx_leaving_atom_flag
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components
 Keywords
Keywords LYASE / PL3 / PARALLEL BETA-HELIX
LYASE / PL3 / PARALLEL BETA-HELIX Function and homology information
Function and homology information pectate lyase /
pectate lyase /  pectate lyase activity / pectin catabolic process /
pectate lyase activity / pectin catabolic process /  hydrolase activity / extracellular region /
hydrolase activity / extracellular region /  membrane /
membrane /  metal ion binding
metal ion binding

 Caldicellulosiruptor bescii (bacteria)
Caldicellulosiruptor bescii (bacteria) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 1.25 Å
SYNCHROTRON / Resolution: 1.25 Å  Authors
Authors United States, 1items
United States, 1items  Citation
Citation Journal: Acta Crystallogr.,Sect.D / Year: 2015
Journal: Acta Crystallogr.,Sect.D / Year: 2015 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 4yzx.cif.gz
4yzx.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb4yzx.ent.gz
pdb4yzx.ent.gz PDB format
PDB format 4yzx.json.gz
4yzx.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/yz/4yzx
https://data.pdbj.org/pub/pdb/validation_reports/yz/4yzx ftp://data.pdbj.org/pub/pdb/validation_reports/yz/4yzx
ftp://data.pdbj.org/pub/pdb/validation_reports/yz/4yzx




 Links
Links Assembly
Assembly


 Components
Components


 Caldicellulosiruptor bescii (strain ATCC BAA-1888 / DSM 6725 / Z-1320) (bacteria)
Caldicellulosiruptor bescii (strain ATCC BAA-1888 / DSM 6725 / Z-1320) (bacteria)
 Escherichia coli (E. coli) / References: UniProt: B9MKT4,
Escherichia coli (E. coli) / References: UniProt: B9MKT4,  pectate lyase
pectate lyase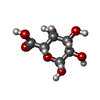
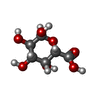

 / Mass: 546.387 Da / Num. of mol.: 2
/ Mass: 546.387 Da / Num. of mol.: 2 D-Galacturonic acid
D-Galacturonic acid D-Galacturonic acid
D-Galacturonic acid










 Imidazole
Imidazole 2-Methyl-2,4-pentanediol
2-Methyl-2,4-pentanediol 2-Methyl-2,4-pentanediol
2-Methyl-2,4-pentanediol Chloride
Chloride Water
Water X-RAY DIFFRACTION
X-RAY DIFFRACTION Sample preparation
Sample preparation
 SYNCHROTRON / Site:
SYNCHROTRON / Site:  SSRL
SSRL  / Beamline: BL11-1 / Wavelength: 0.98397 Å
/ Beamline: BL11-1 / Wavelength: 0.98397 Å : 0.98397 Å / Relative weight: 1
: 0.98397 Å / Relative weight: 1  Processing
Processing Movie
Movie Controller
Controller












 PDBj
PDBj










