[English] 日本語
 Yorodumi
Yorodumi- PDB-4qr9: Crystal structure of two HMGB1 Box A domains cooperating to under... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4qr9 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of two HMGB1 Box A domains cooperating to underwind and kink a DNA | ||||||
 Components Components |
| ||||||
 Keywords Keywords |  DNA BINDING PROTEIN / DNA BINDING PROTEIN /  HMG-box / HMGB1 / Box A domain / HMG-box / HMGB1 / Box A domain /  High mobility group / DNA-binding / bend DNA / kink DNA / architectural factor / High mobility group / DNA-binding / bend DNA / kink DNA / architectural factor /  minor groove / minor groove /  chromatin / chromatin /  Nucleus Nucleus | ||||||
| Function / homology |  Function and homology information Function and homology informationmale-specific defense response to bacterium / positive regulation of myeloid progenitor cell differentiation / open form four-way junction DNA binding / calcium-dependent protein kinase regulator activity / crossed form four-way junction DNA binding / positive regulation of myeloid cell apoptotic process / plasmacytoid dendritic cell activation / regulation of tolerance induction / regulation of T cell mediated immune response to tumor cell / positive regulation of mismatch repair ...male-specific defense response to bacterium / positive regulation of myeloid progenitor cell differentiation / open form four-way junction DNA binding / calcium-dependent protein kinase regulator activity / crossed form four-way junction DNA binding / positive regulation of myeloid cell apoptotic process / plasmacytoid dendritic cell activation / regulation of tolerance induction / regulation of T cell mediated immune response to tumor cell / positive regulation of mismatch repair / negative regulation of apoptotic cell clearance / positive regulation of toll-like receptor 2 signaling pathway / positive regulation of myeloid cell differentiation / negative regulation of RNA polymerase II transcription preinitiation complex assembly / DNA geometric change / myeloid dendritic cell activation / T-helper 1 cell activation / positive regulation of macrophage inflammatory protein 1 alpha production / C-X-C chemokine binding / T-helper 1 cell differentiation / bent DNA binding / positive regulation of glycogen catabolic process /  glycolipid binding / positive regulation of dendritic cell differentiation / negative regulation of CD4-positive, alpha-beta T cell differentiation / positive regulation of toll-like receptor 4 signaling pathway / positive regulation of toll-like receptor 9 signaling pathway / neutrophil clearance / endothelial cell chemotaxis / positive regulation of DNA ligation / glycolipid binding / positive regulation of dendritic cell differentiation / negative regulation of CD4-positive, alpha-beta T cell differentiation / positive regulation of toll-like receptor 4 signaling pathway / positive regulation of toll-like receptor 9 signaling pathway / neutrophil clearance / endothelial cell chemotaxis / positive regulation of DNA ligation /  eye development / positive regulation of interleukin-1 production / eye development / positive regulation of interleukin-1 production /  RAGE receptor binding / induction of positive chemotaxis / alphav-beta3 integrin-HMGB1 complex / bubble DNA binding / RAGE receptor binding / induction of positive chemotaxis / alphav-beta3 integrin-HMGB1 complex / bubble DNA binding /  V(D)J recombination / regulation of nucleotide-excision repair / myeloid cell differentiation / myeloid progenitor cell differentiation / macrophage activation involved in immune response / positive regulation of innate immune response / inflammatory response to antigenic stimulus / positive regulation of monocyte chemotactic protein-1 production / V(D)J recombination / regulation of nucleotide-excision repair / myeloid cell differentiation / myeloid progenitor cell differentiation / macrophage activation involved in immune response / positive regulation of innate immune response / inflammatory response to antigenic stimulus / positive regulation of monocyte chemotactic protein-1 production /  supercoiled DNA binding / cellular response to interleukin-7 / positive regulation of chemokine (C-X-C motif) ligand 2 production / positive regulation of monocyte chemotaxis / glycogen catabolic process / apoptotic cell clearance / myoblast proliferation / endothelial cell proliferation / DNA binding, bending / positive regulation of mesenchymal cell proliferation / positive regulation of vascular endothelial cell proliferation / positive regulation of wound healing / supercoiled DNA binding / cellular response to interleukin-7 / positive regulation of chemokine (C-X-C motif) ligand 2 production / positive regulation of monocyte chemotaxis / glycogen catabolic process / apoptotic cell clearance / myoblast proliferation / endothelial cell proliferation / DNA binding, bending / positive regulation of mesenchymal cell proliferation / positive regulation of vascular endothelial cell proliferation / positive regulation of wound healing /  phosphatidylserine binding / negative regulation of DNA replication / protein kinase activator activity / positive regulation of activated T cell proliferation / positive regulation of sprouting angiogenesis / positive regulation of smooth muscle cell migration / response to type II interferon / positive regulation of interleukin-10 production / negative regulation of blood vessel endothelial cell migration / negative regulation of type II interferon production / phosphatidylserine binding / negative regulation of DNA replication / protein kinase activator activity / positive regulation of activated T cell proliferation / positive regulation of sprouting angiogenesis / positive regulation of smooth muscle cell migration / response to type II interferon / positive regulation of interleukin-10 production / negative regulation of blood vessel endothelial cell migration / negative regulation of type II interferon production /  endoplasmic reticulum-Golgi intermediate compartment / positive regulation of interferon-alpha production / positive regulation of blood vessel endothelial cell migration / positive regulation of myoblast differentiation / cellular response to interleukin-1 / positive regulation of autophagy / response to glucose / heterochromatin formation / endoplasmic reticulum-Golgi intermediate compartment / positive regulation of interferon-alpha production / positive regulation of blood vessel endothelial cell migration / positive regulation of myoblast differentiation / cellular response to interleukin-1 / positive regulation of autophagy / response to glucose / heterochromatin formation /  DNA polymerase binding / four-way junction DNA binding / condensed chromosome / response to glucocorticoid / transcription repressor complex / activation of innate immune response / positive regulation of interleukin-12 production / positive regulation of interferon-beta production / positive regulation of mitotic cell cycle / positive regulation of interleukin-1 beta production / DNA polymerase binding / four-way junction DNA binding / condensed chromosome / response to glucocorticoid / transcription repressor complex / activation of innate immune response / positive regulation of interleukin-12 production / positive regulation of interferon-beta production / positive regulation of mitotic cell cycle / positive regulation of interleukin-1 beta production /  cytokine activity / positive regulation of interleukin-8 production / cytokine activity / positive regulation of interleukin-8 production /  lipopolysaccharide binding / lipopolysaccharide binding /  peptide binding / positive regulation of JNK cascade / lung development / peptide binding / positive regulation of JNK cascade / lung development /  base-excision repair / response to insulin / cell morphogenesis / base-excision repair / response to insulin / cell morphogenesis /  autophagy / positive regulation of neuron projection development / positive regulation of non-canonical NF-kappaB signal transduction / autophagy / positive regulation of neuron projection development / positive regulation of non-canonical NF-kappaB signal transduction /  circadian rhythm / positive regulation of interleukin-6 production / transcription corepressor activity / neuron projection development circadian rhythm / positive regulation of interleukin-6 production / transcription corepressor activity / neuron projection developmentSimilarity search - Function | ||||||
| Biological species |   Rattus norvegicus (Norway rat) Rattus norvegicus (Norway rat)synthetic construct (others) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | ||||||
 Authors Authors | Sanchez-Giraldo, R. / Acosta-Reyes, F.J. / Malarkey, C.S. / Saperas, N. / Churchill, M.E.A. / Campos, J.L. | ||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.D / Year: 2015 Journal: Acta Crystallogr.,Sect.D / Year: 2015Title: Two high-mobility group box domains act together to underwind and kink DNA. Authors: Sanchez-Giraldo, R. / Acosta-Reyes, F.J. / Malarkey, C.S. / Saperas, N. / Churchill, M.E. / Campos, J.L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4qr9.cif.gz 4qr9.cif.gz | 120.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4qr9.ent.gz pdb4qr9.ent.gz | 90.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4qr9.json.gz 4qr9.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qr/4qr9 https://data.pdbj.org/pub/pdb/validation_reports/qr/4qr9 ftp://data.pdbj.org/pub/pdb/validation_reports/qr/4qr9 ftp://data.pdbj.org/pub/pdb/validation_reports/qr/4qr9 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 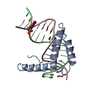 1cktS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||||||||
| Unit cell |
| ||||||||||||||||||
| Noncrystallographic symmetry (NCS) | NCS domain:
NCS domain segments: Component-ID: 0 / Ens-ID: 1 / Beg auth comp-ID: GLY / Beg label comp-ID: GLY / End auth comp-ID: PRO / End label comp-ID: PRO / Refine code: 0 / Auth seq-ID: 5 - 79 / Label seq-ID: 1 - 75
|
- Components
Components
| #1: Protein |  High-mobility group / Amphoterin / Heparin-binding protein p30 / High mobility group protein 1 / HMG-1 High-mobility group / Amphoterin / Heparin-binding protein p30 / High mobility group protein 1 / HMG-1Mass: 8949.302 Da / Num. of mol.: 2 / Fragment: Residues 8-81 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Rattus norvegicus (Norway rat) / Gene: Hmgb1, Hmg-1, Hmg1 / Plasmid: pGEX2T / Production host: Rattus norvegicus (Norway rat) / Gene: Hmgb1, Hmg-1, Hmg1 / Plasmid: pGEX2T / Production host:   Escherichia coli (E. coli) / Strain (production host): Rosetta(DE3)pLySS / References: UniProt: P63159 Escherichia coli (E. coli) / Strain (production host): Rosetta(DE3)pLySS / References: UniProt: P63159#2: DNA chain | Mass: 3043.029 Da / Num. of mol.: 4 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) #3: Chemical | ChemComp-MG / | #4: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.82 Å3/Da / Density % sol: 56.37 % |
|---|---|
Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 6 Details: 0.025M Sodium Cacodylate 0.020M Magnesium Chloride 2.5% MPD (equilibrated against 40% MPD), pH 6, VAPOR DIFFUSION, HANGING DROP, temperature 277K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ALBA ALBA  / Beamline: XALOC / Wavelength: 0.97949 Å / Beamline: XALOC / Wavelength: 0.97949 Å |
| Detector | Type: PSI PILATUS 6M / Detector: PIXEL / Date: Jul 19, 2013 / Details: Mirrors |
| Radiation | Monochromator: Channel-cut Si(111) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.97949 Å / Relative weight: 1 : 0.97949 Å / Relative weight: 1 |
| Reflection | Resolution: 2→42.12 Å / Num. all: 23702 / Num. obs: 23441 / % possible obs: 98.9 % / Observed criterion σ(F): 0 / Observed criterion σ(I): -3 / Redundancy: 7.1 % / Biso Wilson estimate: 32 Å2 / Rmerge(I) obs: 0.113 / Net I/σ(I): 16.025 |
| Reflection shell | Resolution: 2→2.07 Å / Redundancy: 5.1 % / Rmerge(I) obs: 0.658 / Mean I/σ(I) obs: 1.9 / Num. unique all: 2162 / % possible all: 93.5 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB entry 1CKT Resolution: 2→42.12 Å / Cor.coef. Fo:Fc: 0.949 / Cor.coef. Fo:Fc free: 0.941 / SU B: 6.442 / SU ML: 0.097 / Cross valid method: THROUGHOUT / ESU R: 0.161 / ESU R Free: 0.15 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 42.412 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→42.12 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS | Ens-ID: 1 / Number: 3839 / Refine-ID: X-RAY DIFFRACTION / Type: interatomic distance / Rms dev position: 0.17 Å / Weight position: 0.05
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.001→2.053 Å / Total num. of bins used: 20
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj





