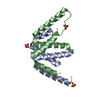
Entry Database : PDB / ID : 4myzTitle Structure of a class 2 docking domain complex from modules CurK and CurL of the curacin A polyketide synthase CurK, CurL fusion protein Keywords / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Moorea producens 3L (bacteria)Method / / / Resolution : 1.5 Å Authors Whicher, J.R. / Smaga, S.S. / Smith, J.L. Journal : Chem.Biol. / Year : 2013Title : Cyanobacterial polyketide synthase docking domains: a tool for engineering natural product biosynthesis.Authors : Whicher, J.R. / Smaga, S.S. / Hansen, D.A. / Brown, W.C. / Gerwick, W.H. / Sherman, D.H. / Smith, J.L. History Deposition Sep 28, 2013 Deposition site / Processing site Revision 1.0 Jan 29, 2014 Provider / Type Revision 1.1 Feb 5, 2014 Group Revision 1.2 Aug 9, 2017 Group / Refinement description / Source and taxonomyCategory / entity_src_gen / software / Item Revision 1.3 Feb 28, 2024 Group / Database referencesCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / struct_ref_seq_dif Item / _database_2.pdbx_database_accession / _struct_ref_seq_dif.details
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords PROTEIN BINDING /
PROTEIN BINDING /  Protein-protein interaction /
Protein-protein interaction /  fusion protein
fusion protein Function and homology information
Function and homology information : / S-adenosylmethionine-dependent methyltransferase activity /
: / S-adenosylmethionine-dependent methyltransferase activity /  acyltransferase activity /
acyltransferase activity /  phosphopantetheine binding / 3-oxoacyl-[acyl-carrier-protein] synthase activity / fatty acid biosynthetic process /
phosphopantetheine binding / 3-oxoacyl-[acyl-carrier-protein] synthase activity / fatty acid biosynthetic process /  oxidoreductase activity /
oxidoreductase activity /  metal ion binding
metal ion binding Moorea producens 3L (bacteria)
Moorea producens 3L (bacteria) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.5 Å
MOLECULAR REPLACEMENT / Resolution: 1.5 Å  Authors
Authors Citation
Citation Journal: Chem.Biol. / Year: 2013
Journal: Chem.Biol. / Year: 2013 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 4myz.cif.gz
4myz.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb4myz.ent.gz
pdb4myz.ent.gz PDB format
PDB format 4myz.json.gz
4myz.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/my/4myz
https://data.pdbj.org/pub/pdb/validation_reports/my/4myz ftp://data.pdbj.org/pub/pdb/validation_reports/my/4myz
ftp://data.pdbj.org/pub/pdb/validation_reports/my/4myz Links
Links Assembly
Assembly



 Components
Components Moorea producens 3L (bacteria) / Gene: LYNGBM3L_74450, LYNGBM3L_74440 / Plasmid: pMCSG7 / Production host:
Moorea producens 3L (bacteria) / Gene: LYNGBM3L_74450, LYNGBM3L_74440 / Plasmid: pMCSG7 / Production host: 
 Escherichia coli (E. coli) / Strain (production host): BL21(DE3) / References: UniProt: F4Y425, UniProt: F4Y424
Escherichia coli (E. coli) / Strain (production host): BL21(DE3) / References: UniProt: F4Y425, UniProt: F4Y424 Water
Water X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation
 SYNCHROTRON / Site:
SYNCHROTRON / Site:  APS
APS  / Beamline: 23-ID-D / Wavelength: 1.033 Å
/ Beamline: 23-ID-D / Wavelength: 1.033 Å : 1.033 Å / Relative weight: 1
: 1.033 Å / Relative weight: 1  Processing
Processing :
:  MOLECULAR REPLACEMENT / Resolution: 1.5→26.45 Å / Cor.coef. Fo:Fc: 0.961 / Cor.coef. Fo:Fc free: 0.954 / SU B: 2.106 / SU ML: 0.038 / Cross valid method: THROUGHOUT / ESU R: 0.06 / ESU R Free: 0.063 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
MOLECULAR REPLACEMENT / Resolution: 1.5→26.45 Å / Cor.coef. Fo:Fc: 0.961 / Cor.coef. Fo:Fc free: 0.954 / SU B: 2.106 / SU ML: 0.038 / Cross valid method: THROUGHOUT / ESU R: 0.06 / ESU R Free: 0.063 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS Movie
Movie Controller
Controller













 PDBj
PDBj




