+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4jnb | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of the Catalytic Domain of Human DUSP12 | ||||||
 Components Components | Dual specificity protein phosphatase 12 | ||||||
 Keywords Keywords |  HYDROLASE / HYDROLASE /  DUSP / Dual specificity phosphatase / DUSP / Dual specificity phosphatase /  phosphatase phosphatase | ||||||
| Function / homology |  Function and homology information Function and homology informationprotein tyrosine/serine/threonine phosphatase activity / myosin phosphatase activity / protein-serine/threonine phosphatase /  phosphatase activity / phosphatase activity /  dephosphorylation / dephosphorylation /  protein-tyrosine-phosphatase / protein-tyrosine-phosphatase /  protein tyrosine phosphatase activity / protein modification process / protein tyrosine phosphatase activity / protein modification process /  kinase binding / zinc ion binding ...protein tyrosine/serine/threonine phosphatase activity / myosin phosphatase activity / protein-serine/threonine phosphatase / kinase binding / zinc ion binding ...protein tyrosine/serine/threonine phosphatase activity / myosin phosphatase activity / protein-serine/threonine phosphatase /  phosphatase activity / phosphatase activity /  dephosphorylation / dephosphorylation /  protein-tyrosine-phosphatase / protein-tyrosine-phosphatase /  protein tyrosine phosphatase activity / protein modification process / protein tyrosine phosphatase activity / protein modification process /  kinase binding / zinc ion binding / kinase binding / zinc ion binding /  nucleoplasm / nucleoplasm /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 3 Å MOLECULAR REPLACEMENT / Resolution: 3 Å | ||||||
 Authors Authors | Jeon, T.J. / Chien, P.N. / Ku, B. / Kim, S.J. / Ryu, S.E. | ||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.D / Year: 2014 Journal: Acta Crystallogr.,Sect.D / Year: 2014Title: The family-wide structure and function of human dual-specificity protein phosphatases. Authors: Jeong, D.G. / Wei, C.H. / Ku, B. / Jeon, T.J. / Chien, P.N. / Kim, J.K. / Park, S.Y. / Hwang, H.S. / Ryu, S.Y. / Park, H. / Kim, D.S. / Kim, S.J. / Ryu, S.E. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4jnb.cif.gz 4jnb.cif.gz | 42.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4jnb.ent.gz pdb4jnb.ent.gz | 29.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4jnb.json.gz 4jnb.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/jn/4jnb https://data.pdbj.org/pub/pdb/validation_reports/jn/4jnb ftp://data.pdbj.org/pub/pdb/validation_reports/jn/4jnb ftp://data.pdbj.org/pub/pdb/validation_reports/jn/4jnb | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 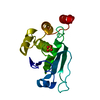
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 18619.234 Da / Num. of mol.: 1 / Fragment: catalytic domain UNP RESIDUES 27-193 / Mutation: C97A, C115S Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: DUSP12 / Production host: Homo sapiens (human) / Gene: DUSP12 / Production host:   Escherichia coli (E. coli) Escherichia coli (E. coli)References: UniProt: Q9UNI6, protein-serine/threonine phosphatase,  protein-tyrosine-phosphatase protein-tyrosine-phosphatase |
|---|---|
| #2: Chemical | ChemComp-SO4 /  Sulfate Sulfate |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.38 Å3/Da / Density % sol: 63.58 % |
|---|---|
Crystal grow | Temperature: 298 K / Method: vapor diffusion / pH: 8.5 Details: 0.1M Tris pH 8.5, 1.9M Ammonium sulfate, VAPOR DIFFUSION, temperature 298K |
-Data collection
| Diffraction source | Source:  SYNCHROTRON / Site: PAL/PLS SYNCHROTRON / Site: PAL/PLS  / Beamline: 4A / Beamline: 4A |
|---|---|
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Relative weight: 1 |
| Reflection | Highest resolution: 3 Å / Num. obs: 4743 |
- Processing
Processing
| Software | Name: CNS / Classification: refinement | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT / Resolution: 3→50 Å MOLECULAR REPLACEMENT / Resolution: 3→50 Å
| |||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 3→50 Å
|
 Movie
Movie Controller
Controller














 PDBj
PDBj








