+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4i1k | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal Structure of VRN1 (Residues 208-341) | ||||||
 Components Components | B3 domain-containing transcription factor VRN1 | ||||||
 Keywords Keywords |  DNA BINDING PROTEIN / B3 domain beta-barrel DNA BINDING PROTEIN / B3 domain beta-barrel | ||||||
| Function / homology |  Function and homology information Function and homology informationvernalization response /  regulation of flower development / transcription cis-regulatory region binding / regulation of flower development / transcription cis-regulatory region binding /  DNA binding / DNA binding /  nucleoplasm nucleoplasmSimilarity search - Function | ||||||
| Biological species |   Arabidopsis thaliana (thale cress) Arabidopsis thaliana (thale cress) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SAD / Resolution: 1.6 Å SAD / Resolution: 1.6 Å | ||||||
 Authors Authors | King, G. / Chanson, A.H. / McCallum, E.J. / Ohme-Takagi, M. / Byriel, K. / Hill, J.M. / Martin, J.L. / Mylne, J.S. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2013 Journal: J.Biol.Chem. / Year: 2013Title: The Arabidopsis B3 Domain Protein VERNALIZATION1 (VRN1) Is Involved in Processes Essential for Development, with Structural and Mutational Studies Revealing Its DNA-binding Surface. Authors: King, G.J. / Chanson, A.H. / McCallum, E.J. / Ohme-Takagi, M. / Byriel, K. / Hill, J.M. / Martin, J.L. / Mylne, J.S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4i1k.cif.gz 4i1k.cif.gz | 125.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4i1k.ent.gz pdb4i1k.ent.gz | 98.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4i1k.json.gz 4i1k.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/i1/4i1k https://data.pdbj.org/pub/pdb/validation_reports/i1/4i1k ftp://data.pdbj.org/pub/pdb/validation_reports/i1/4i1k ftp://data.pdbj.org/pub/pdb/validation_reports/i1/4i1k | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 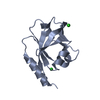
| |||||||||
| 2 | 
| |||||||||
| Unit cell |
| |||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 17017.479 Da / Num. of mol.: 2 / Fragment: B3 domain (UNP residues 208-341) Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Arabidopsis thaliana (thale cress) / Gene: VRN1, At3g18990, K13E13.10 / Production host: Arabidopsis thaliana (thale cress) / Gene: VRN1, At3g18990, K13E13.10 / Production host:   Escherichia coli (E. coli) / References: UniProt: Q8L3W1 Escherichia coli (E. coli) / References: UniProt: Q8L3W1#2: Chemical | ChemComp-CL /  Chloride Chloride#3: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 2 X-RAY DIFFRACTION / Number of used crystals: 2 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.05 Å3/Da / Density % sol: 40.11 % |
|---|---|
Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 6 Details: 19 mg/mL protein in 10 mM HEPES, pH 7.0, 50 mM sodium chloride, 5 mM DTT + equal volume well solution (0.1 M MES, pH 6.0, 1.4 M sodium chloride, 10 mM manganese chloride), VAPOR DIFFUSION, ...Details: 19 mg/mL protein in 10 mM HEPES, pH 7.0, 50 mM sodium chloride, 5 mM DTT + equal volume well solution (0.1 M MES, pH 6.0, 1.4 M sodium chloride, 10 mM manganese chloride), VAPOR DIFFUSION, HANGING DROP, temperature 277K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Australian Synchrotron Australian Synchrotron  / Beamline: MX1 / Wavelength: 0.956 Å / Beamline: MX1 / Wavelength: 0.956 Å |
| Detector | Type: ADSC QUANTUM 210r / Detector: CCD / Date: Aug 16, 2008 |
| Radiation | Monochromator: double crystal Si(111) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.956 Å / Relative weight: 1 : 0.956 Å / Relative weight: 1 |
| Reflection | Resolution: 1.6→50 Å / Num. all: 34680 / Num. obs: 34680 / % possible obs: 100 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 |
- Processing
Processing
| Software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  SAD / Resolution: 1.6→50 Å / σ(F): 1 / Stereochemistry target values: Engh & Huber SAD / Resolution: 1.6→50 Å / σ(F): 1 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.6→50 Å
| ||||||||||||||||||||
| LS refinement shell | Resolution: 1.6→1.66 Å / Num. reflection obs: 3439 |
 Movie
Movie Controller
Controller













 PDBj
PDBj


