+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3jve | ||||||
|---|---|---|---|---|---|---|---|
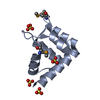
| Title | Crystal Structure of the Sixth BRCT Domain of TopBP1 | ||||||
 Components Components | DNA topoisomerase 2-binding protein 1 | ||||||
 Keywords Keywords |  PROTEIN BINDING / PROTEIN BINDING /  BRCT domain / BRCT domain /  DNA damage / DNA damage /  DNA repair / DNA-binding / DNA repair / DNA-binding /  Nucleus / Nucleus /  Phosphoprotein / Polymorphism / Ubl conjugation / Phosphoprotein / Polymorphism / Ubl conjugation /  DNA BINDING PROTEIN DNA BINDING PROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology informationbroken chromosome clustering / BRCA1-B complex / phosphorylation-dependent protein binding / chromatin-protein adaptor activity /  homologous recombination / protein localization to site of double-strand break / DNA replication checkpoint signaling / double-strand break repair via classical nonhomologous end joining / mitotic DNA replication checkpoint signaling / double-strand break repair via alternative nonhomologous end joining ...broken chromosome clustering / BRCA1-B complex / phosphorylation-dependent protein binding / chromatin-protein adaptor activity / homologous recombination / protein localization to site of double-strand break / DNA replication checkpoint signaling / double-strand break repair via classical nonhomologous end joining / mitotic DNA replication checkpoint signaling / double-strand break repair via alternative nonhomologous end joining ...broken chromosome clustering / BRCA1-B complex / phosphorylation-dependent protein binding / chromatin-protein adaptor activity /  homologous recombination / protein localization to site of double-strand break / DNA replication checkpoint signaling / double-strand break repair via classical nonhomologous end joining / mitotic DNA replication checkpoint signaling / double-strand break repair via alternative nonhomologous end joining / HDR through Single Strand Annealing (SSA) / Impaired BRCA2 binding to RAD51 / DNA metabolic process / response to ionizing radiation / mitotic G2 DNA damage checkpoint signaling / site of DNA damage / Presynaptic phase of homologous DNA pairing and strand exchange / chromosome organization / DNA replication initiation / protein serine/threonine kinase activator activity / DNA damage checkpoint signaling / condensed nuclear chromosome / male germ cell nucleus / double-strand break repair via homologous recombination / G2/M DNA damage checkpoint / PML body / homologous recombination / protein localization to site of double-strand break / DNA replication checkpoint signaling / double-strand break repair via classical nonhomologous end joining / mitotic DNA replication checkpoint signaling / double-strand break repair via alternative nonhomologous end joining / HDR through Single Strand Annealing (SSA) / Impaired BRCA2 binding to RAD51 / DNA metabolic process / response to ionizing radiation / mitotic G2 DNA damage checkpoint signaling / site of DNA damage / Presynaptic phase of homologous DNA pairing and strand exchange / chromosome organization / DNA replication initiation / protein serine/threonine kinase activator activity / DNA damage checkpoint signaling / condensed nuclear chromosome / male germ cell nucleus / double-strand break repair via homologous recombination / G2/M DNA damage checkpoint / PML body /  spindle pole / spindle pole /  actin cytoskeleton / site of double-strand break / actin cytoskeleton / site of double-strand break /  chromosome / Processing of DNA double-strand break ends / Regulation of TP53 Activity through Phosphorylation / chromosome / Processing of DNA double-strand break ends / Regulation of TP53 Activity through Phosphorylation /  nuclear body / nuclear body /  DNA repair / DNA repair /  centrosome / DNA damage response / centrosome / DNA damage response /  DNA binding / DNA binding /  nucleoplasm / identical protein binding / nucleoplasm / identical protein binding /  nucleus / nucleus /  plasma membrane / plasma membrane /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 1.34 Å molecular replacement / Resolution: 1.34 Å | ||||||
 Authors Authors | Leung, C.C. / Kellogg, E. / Baker, D. / Glover, J.N.M. | ||||||
 Citation Citation |  Journal: Protein Sci. / Year: 2010 Journal: Protein Sci. / Year: 2010Title: Insights from the crystal structure of the sixth BRCT domain of topoisomerase IIbeta binding protein 1. Authors: Leung, C.C. / Kellogg, E. / Kuhnert, A. / Hanel, F. / Baker, D. / Glover, J.N. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3jve.cif.gz 3jve.cif.gz | 58.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3jve.ent.gz pdb3jve.ent.gz | 42.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3jve.json.gz 3jve.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/jv/3jve https://data.pdbj.org/pub/pdb/validation_reports/jv/3jve ftp://data.pdbj.org/pub/pdb/validation_reports/jv/3jve ftp://data.pdbj.org/pub/pdb/validation_reports/jv/3jve | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 12412.957 Da / Num. of mol.: 1 / Fragment: C-terminus (BRCT) domain Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: TOPBP1, KIAA0259 / Production host: Homo sapiens (human) / Gene: TOPBP1, KIAA0259 / Production host:   Escherichia coli (E. coli) / References: UniProt: Q92547 Escherichia coli (E. coli) / References: UniProt: Q92547 |
|---|---|
| #2: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.32 Å3/Da / Density % sol: 46.93 % |
|---|---|
Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 6.8 Details: 0.1M Tris-HCl pH 6.8, PEG 2000 MME, VAPOR DIFFUSION, HANGING DROP, temperature 277K |
-Data collection
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  CLSI CLSI  / Beamline: 08ID-1 / Wavelength: 0.97934 Å / Beamline: 08ID-1 / Wavelength: 0.97934 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Type: MARMOSAIC 225 mm CCD / Detector: CCD / Date: Jul 1, 2008 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength : 0.97934 Å / Relative weight: 1 : 0.97934 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 1.34→50 Å / Num. obs: 26433 / % possible obs: 99 % / Redundancy: 6.5 % / Rmerge(I) obs: 0.054 / Χ2: 0.764 / Net I/σ(I): 9 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell |
|
-Phasing
Phasing | Method:  molecular replacement molecular replacement | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Phasing MR | Rfactor: 43.85 / Model details: Phaser MODE: MR_AUTO
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT / Resolution: 1.34→39.78 Å / Cor.coef. Fo:Fc: 0.968 / Cor.coef. Fo:Fc free: 0.961 / WRfactor Rfree: 0.173 / WRfactor Rwork: 0.158 / Occupancy max: 1 / Occupancy min: 0.2 / FOM work R set: 0.919 / SU B: 1.222 / SU ML: 0.024 / SU R Cruickshank DPI: 0.053 / SU Rfree: 0.047 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.053 / ESU R Free: 0.047 / Stereochemistry target values: MAXIMUM LIKELIHOOD MOLECULAR REPLACEMENT / Resolution: 1.34→39.78 Å / Cor.coef. Fo:Fc: 0.968 / Cor.coef. Fo:Fc free: 0.961 / WRfactor Rfree: 0.173 / WRfactor Rwork: 0.158 / Occupancy max: 1 / Occupancy min: 0.2 / FOM work R set: 0.919 / SU B: 1.222 / SU ML: 0.024 / SU R Cruickshank DPI: 0.053 / SU Rfree: 0.047 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.053 / ESU R Free: 0.047 / Stereochemistry target values: MAXIMUM LIKELIHOODDetails: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS U VALUES : RESIDUAL ONLY
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 47.13 Å2 / Biso mean: 12.118 Å2 / Biso min: 4.51 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.34→39.78 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.339→1.374 Å / Total num. of bins used: 20
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 9.378 Å / Origin y: 72.477 Å / Origin z: 10.097 Å
|
 Movie
Movie Controller
Controller











 PDBj
PDBj




