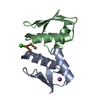
Entry Database : PDB / ID : 2xmuTitle Copper chaperone Atx1 from Synechocystis PCC6803 (Cu2 form) SSR2857 PROTEIN Keywords / / / / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / Biological species SYNECHOCYSTIS SP. PCC 6803 (bacteria)Method / / / Resolution : 1.75 Å Authors Badarau, A. / Firbank, S.J. / McCarthy, A.A. / Banfield, M.J. / Dennison, C. Journal : Biochemistry / Year : 2010Title : Visualizing the Metal-Binding Versatility of Copper Trafficking Sites .Authors : Badarau, A. / Firbank, S.J. / Mccarthy, A.A. / Banfield, M.J. / Dennison, C. History Deposition Jul 29, 2010 Deposition site / Processing site Revision 1.0 Aug 18, 2010 Provider / Type Revision 1.1 Dec 3, 2014 Group / Version format complianceRevision 1.2 Dec 20, 2023 Group Data collection / Database references ... Data collection / Database references / Derived calculations / Other / Refinement description Category chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_database_status / pdbx_initial_refinement_model / pdbx_struct_conn_angle / struct_conn / struct_site Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_database_status.status_code_sf / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords CHAPERONE / COPPER HOMEOSTASIS / P-TYPE ATPASES / METAL TRANSPORT / METALLOCHAPERONE / CU(I)-BINDING / CU(I)-CLUSTER / TRAFFICKING
CHAPERONE / COPPER HOMEOSTASIS / P-TYPE ATPASES / METAL TRANSPORT / METALLOCHAPERONE / CU(I)-BINDING / CU(I)-CLUSTER / TRAFFICKING Function and homology information
Function and homology information
 SYNECHOCYSTIS SP. PCC 6803 (bacteria)
SYNECHOCYSTIS SP. PCC 6803 (bacteria) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.75 Å
MOLECULAR REPLACEMENT / Resolution: 1.75 Å  Authors
Authors Citation
Citation Journal: Biochemistry / Year: 2010
Journal: Biochemistry / Year: 2010 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2xmu.cif.gz
2xmu.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2xmu.ent.gz
pdb2xmu.ent.gz PDB format
PDB format 2xmu.json.gz
2xmu.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/xm/2xmu
https://data.pdbj.org/pub/pdb/validation_reports/xm/2xmu ftp://data.pdbj.org/pub/pdb/validation_reports/xm/2xmu
ftp://data.pdbj.org/pub/pdb/validation_reports/xm/2xmu




 Links
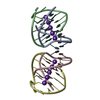
Links Assembly
Assembly
 Components
Components
 SYNECHOCYSTIS SP. PCC 6803 (bacteria) / Plasmid: PET29A / Production host:
SYNECHOCYSTIS SP. PCC 6803 (bacteria) / Plasmid: PET29A / Production host: 
 ESCHERICHIA COLI (E. coli) / Strain (production host): BL21(DE3) / References: UniProt: P73213
ESCHERICHIA COLI (E. coli) / Strain (production host): BL21(DE3) / References: UniProt: P73213 Copper
Copper Water
Water X-RAY DIFFRACTION / Number of used crystals: 2
X-RAY DIFFRACTION / Number of used crystals: 2  Sample preparation
Sample preparation
 : 1.38 Å / Relative weight: 1
: 1.38 Å / Relative weight: 1  Processing
Processing :
:  MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller











 PDBj
PDBj



