[English] 日本語
 Yorodumi
Yorodumi- PDB-2rr0: Structure of epidermal growth factor-like repeat 12 of mouse Notc... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2rr0 | ||||||
|---|---|---|---|---|---|---|---|
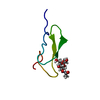
| Title | Structure of epidermal growth factor-like repeat 12 of mouse Notch-1 receptor | ||||||
 Components Components | Neurogenic locus notch homolog protein 1 | ||||||
 Keywords Keywords |  RECEPTOR / Notch / RECEPTOR / Notch /  EGF-Like Domain / Activator / ANK repeat / EGF-Like Domain / Activator / ANK repeat /  Cell membrane / Cell membrane /  Developmental protein / Differentiation / Developmental protein / Differentiation /  Disulfide bond / Disulfide bond /  Glycoprotein / Metal-binding / Glycoprotein / Metal-binding /  Notch signaling pathway / Notch signaling pathway /  Nucleus / Nucleus /  Phosphoprotein / Phosphoprotein /  Transcription / Transcription /  Transcription regulation / Transcription regulation /  Transmembrane Transmembrane | ||||||
| Function / homology |  Function and homology information Function and homology informationPre-NOTCH Processing in Golgi / regulation of cardioblast proliferation / regulation of inner ear auditory receptor cell differentiation / positive regulation of ephrin receptor signaling pathway / positive regulation of glial cell differentiation / osteoblast fate commitment / venous blood vessel morphogenesis / Activated NOTCH1 Transmits Signal to the Nucleus / coronary sinus valve morphogenesis / cardiac right atrium morphogenesis ...Pre-NOTCH Processing in Golgi / regulation of cardioblast proliferation / regulation of inner ear auditory receptor cell differentiation / positive regulation of ephrin receptor signaling pathway / positive regulation of glial cell differentiation / osteoblast fate commitment / venous blood vessel morphogenesis / Activated NOTCH1 Transmits Signal to the Nucleus / coronary sinus valve morphogenesis / cardiac right atrium morphogenesis / cardiac right ventricle formation / growth involved in heart morphogenesis / Notch signaling pathway involved in regulation of secondary heart field cardioblast proliferation /  cell differentiation in spinal cord / retinal cone cell differentiation / venous endothelial cell differentiation / arterial endothelial cell differentiation / cardiac chamber formation / epithelial cell fate commitment / negative regulation of pro-B cell differentiation / negative regulation of inner ear auditory receptor cell differentiation / mitral valve formation / cell migration involved in endocardial cushion formation / glomerular mesangial cell development / negative regulation of photoreceptor cell differentiation / negative regulation of cell proliferation involved in heart valve morphogenesis / cell differentiation in spinal cord / retinal cone cell differentiation / venous endothelial cell differentiation / arterial endothelial cell differentiation / cardiac chamber formation / epithelial cell fate commitment / negative regulation of pro-B cell differentiation / negative regulation of inner ear auditory receptor cell differentiation / mitral valve formation / cell migration involved in endocardial cushion formation / glomerular mesangial cell development / negative regulation of photoreceptor cell differentiation / negative regulation of cell proliferation involved in heart valve morphogenesis /  regulation of somitogenesis / inhibition of neuroepithelial cell differentiation / endocardium morphogenesis / atrioventricular node development / foregut morphogenesis / regulation of cell adhesion involved in heart morphogenesis / distal tubule development / MAML1-RBP-Jkappa- ICN1 complex / regulation of epithelial cell proliferation involved in prostate gland development / auditory receptor cell fate commitment / positive regulation of aorta morphogenesis / negative regulation of endothelial cell chemotaxis / NOTCH1 Intracellular Domain Regulates Transcription / RUNX3 regulates NOTCH signaling / neuroendocrine cell differentiation / collecting duct development / negative regulation of extracellular matrix constituent secretion / positive regulation of transcription of Notch receptor target / cellular response to tumor cell / positive regulation of apoptotic process involved in morphogenesis / Notch-HLH transcription pathway / compartment pattern specification / vasculogenesis involved in coronary vascular morphogenesis / T-helper 17 type immune response / endocardial cushion development / regulation of somitogenesis / inhibition of neuroepithelial cell differentiation / endocardium morphogenesis / atrioventricular node development / foregut morphogenesis / regulation of cell adhesion involved in heart morphogenesis / distal tubule development / MAML1-RBP-Jkappa- ICN1 complex / regulation of epithelial cell proliferation involved in prostate gland development / auditory receptor cell fate commitment / positive regulation of aorta morphogenesis / negative regulation of endothelial cell chemotaxis / NOTCH1 Intracellular Domain Regulates Transcription / RUNX3 regulates NOTCH signaling / neuroendocrine cell differentiation / collecting duct development / negative regulation of extracellular matrix constituent secretion / positive regulation of transcription of Notch receptor target / cellular response to tumor cell / positive regulation of apoptotic process involved in morphogenesis / Notch-HLH transcription pathway / compartment pattern specification / vasculogenesis involved in coronary vascular morphogenesis / T-helper 17 type immune response / endocardial cushion development /  regulation of extracellular matrix assembly / endocardial cell differentiation / epithelial to mesenchymal transition involved in endocardial cushion formation / cardiac ventricle morphogenesis / positive regulation of smooth muscle cell differentiation / cardiac left ventricle morphogenesis / mesenchymal cell development / epidermal cell fate specification / regulation of extracellular matrix assembly / endocardial cell differentiation / epithelial to mesenchymal transition involved in endocardial cushion formation / cardiac ventricle morphogenesis / positive regulation of smooth muscle cell differentiation / cardiac left ventricle morphogenesis / mesenchymal cell development / epidermal cell fate specification /  regulation of Notch signaling pathway / coronary vein morphogenesis / negative regulation of collagen biosynthetic process / cardiac vascular smooth muscle cell development / negative regulation of myotube differentiation / somatic stem cell division / left/right axis specification / negative regulation of cardiac muscle hypertrophy / negative regulation of cell adhesion molecule production / interleukin-17-mediated signaling pathway / positive regulation of endothelial cell differentiation / secretory columnal luminar epithelial cell differentiation involved in prostate glandular acinus development / endocardium development / apoptotic process involved in embryonic digit morphogenesis / glial cell differentiation / positive regulation of cardiac epithelial to mesenchymal transition / cardiac epithelial to mesenchymal transition / negative regulation of calcium ion-dependent exocytosis / cardiac muscle cell myoblast differentiation / cellular response to follicle-stimulating hormone stimulus / neuron fate commitment / pericardium morphogenesis / cardiac atrium morphogenesis / negative regulation of catalytic activity / neuronal stem cell population maintenance / regulation of Notch signaling pathway / coronary vein morphogenesis / negative regulation of collagen biosynthetic process / cardiac vascular smooth muscle cell development / negative regulation of myotube differentiation / somatic stem cell division / left/right axis specification / negative regulation of cardiac muscle hypertrophy / negative regulation of cell adhesion molecule production / interleukin-17-mediated signaling pathway / positive regulation of endothelial cell differentiation / secretory columnal luminar epithelial cell differentiation involved in prostate glandular acinus development / endocardium development / apoptotic process involved in embryonic digit morphogenesis / glial cell differentiation / positive regulation of cardiac epithelial to mesenchymal transition / cardiac epithelial to mesenchymal transition / negative regulation of calcium ion-dependent exocytosis / cardiac muscle cell myoblast differentiation / cellular response to follicle-stimulating hormone stimulus / neuron fate commitment / pericardium morphogenesis / cardiac atrium morphogenesis / negative regulation of catalytic activity / neuronal stem cell population maintenance /  tissue regeneration / regulation of stem cell proliferation / negative regulation of oligodendrocyte differentiation / positive regulation of astrocyte differentiation / calcium-ion regulated exocytosis / pulmonary valve morphogenesis / heart trabecula morphogenesis / negative regulation of biomineral tissue development / endoderm development / coronary artery morphogenesis / negative regulation of cell-cell adhesion mediated by cadherin / prostate gland epithelium morphogenesis / tissue regeneration / regulation of stem cell proliferation / negative regulation of oligodendrocyte differentiation / positive regulation of astrocyte differentiation / calcium-ion regulated exocytosis / pulmonary valve morphogenesis / heart trabecula morphogenesis / negative regulation of biomineral tissue development / endoderm development / coronary artery morphogenesis / negative regulation of cell-cell adhesion mediated by cadherin / prostate gland epithelium morphogenesis /  luteolysis / cardiac muscle tissue morphogenesis / ventricular trabecula myocardium morphogenesis / negative regulation of myoblast differentiation luteolysis / cardiac muscle tissue morphogenesis / ventricular trabecula myocardium morphogenesis / negative regulation of myoblast differentiationSimilarity search - Function | ||||||
| Biological species |   Mus musculus (house mouse) Mus musculus (house mouse) | ||||||
| Method |  SOLUTION NMR / DGSA-distance geometry simulated annealing SOLUTION NMR / DGSA-distance geometry simulated annealing | ||||||
| Model details | lowest energy, model 1 | ||||||
 Authors Authors | Hosoguchi, K. / Shimizu, K. / Fujitani, N. / Nishimura, S. | ||||||
 Citation Citation |  Journal: J.Am.Chem.Soc. / Year: 2010 Journal: J.Am.Chem.Soc. / Year: 2010Title: Chemical Synthesis, Folding, and Structural Insights into O-Fucosylated Epidermal Growth Factor-like Repeat 12 of Mouse Notch-1 Receptor Authors: Hiruma-Shimizu, K. / Hosoguchi, K. / Liu, Y. / Fujitani, N. / Ohta, T. / Hinou, H. / Matsushita, T. / Shimizu, H. / Feizi, T. / Nishimura, S. | ||||||
| History |
| ||||||
| Remark 700 | SHEET DETERMINATION METHOD: AUTHOR DETERMINED |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2rr0.cif.gz 2rr0.cif.gz | 212.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2rr0.ent.gz pdb2rr0.ent.gz | 183.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2rr0.json.gz 2rr0.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/rr/2rr0 https://data.pdbj.org/pub/pdb/validation_reports/rr/2rr0 ftp://data.pdbj.org/pub/pdb/validation_reports/rr/2rr0 ftp://data.pdbj.org/pub/pdb/validation_reports/rr/2rr0 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 4218.697 Da / Num. of mol.: 1 / Fragment: EXTRACELLULAR DOMAIN EGF-LIKE repeat 12 / Source method: obtained synthetically / Details: synthetic peptide / Source: (synth.)   Mus musculus (house mouse) / References: UniProt: Q01705 Mus musculus (house mouse) / References: UniProt: Q01705 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||
| NMR details | Text: The structure was determined using a combination NOE and D2O exchange experiment. |
- Sample preparation
Sample preparation
| Details |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| |||||||||
| Sample conditions | Ionic strength: 0 / pH: 5.3 / Pressure: ambient / Temperature: 300 K |
-NMR measurement
| NMR spectrometer | Type: Bruker Avance / Manufacturer: Bruker / Model : Avance / Field strength: 600 MHz : Avance / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software | Name: CNS / Version: 1.1 / Classification: refinement |
|---|---|
| Refinement | Method: DGSA-distance geometry simulated annealing / Software ordinal: 1 |
| NMR representative | Selection criteria: lowest energy |
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 200 / Conformers submitted total number: 20 / Representative conformer: 1 |
 Movie
Movie Controller
Controller













 PDBj
PDBj







