+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2ms8 | ||||||
|---|---|---|---|---|---|---|---|
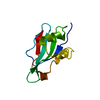
| Title | Solution NMR structure of MAVS CARD | ||||||
 Components Components | Mitochondrial antiviral-signaling protein | ||||||
 Keywords Keywords |  PROTEIN BINDING / MAVS CARD PROTEIN BINDING / MAVS CARD | ||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of IP-10 production / regulation of peroxisome organization / RIG-I binding / positive regulation of chemokine (C-C motif) ligand 5 production / positive regulation of myeloid dendritic cell cytokine production /  CARD domain binding / NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 / protein localization to mitochondrion / positive regulation of response to cytokine stimulus / positive regulation of type I interferon-mediated signaling pathway ...positive regulation of IP-10 production / regulation of peroxisome organization / RIG-I binding / positive regulation of chemokine (C-C motif) ligand 5 production / positive regulation of myeloid dendritic cell cytokine production / CARD domain binding / NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 / protein localization to mitochondrion / positive regulation of response to cytokine stimulus / positive regulation of type I interferon-mediated signaling pathway ...positive regulation of IP-10 production / regulation of peroxisome organization / RIG-I binding / positive regulation of chemokine (C-C motif) ligand 5 production / positive regulation of myeloid dendritic cell cytokine production /  CARD domain binding / NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 / protein localization to mitochondrion / positive regulation of response to cytokine stimulus / positive regulation of type I interferon-mediated signaling pathway / peroxisomal membrane / TRAF6 mediated IRF7 activation / negative regulation of type I interferon-mediated signaling pathway / positive regulation of NLRP3 inflammasome complex assembly / negative regulation of viral genome replication / type I interferon-mediated signaling pathway / cellular response to exogenous dsRNA / cytoplasmic pattern recognition receptor signaling pathway / positive regulation of interferon-alpha production / antiviral innate immune response / TRAF6 mediated NF-kB activation / positive regulation of type I interferon production / CARD domain binding / NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 / protein localization to mitochondrion / positive regulation of response to cytokine stimulus / positive regulation of type I interferon-mediated signaling pathway / peroxisomal membrane / TRAF6 mediated IRF7 activation / negative regulation of type I interferon-mediated signaling pathway / positive regulation of NLRP3 inflammasome complex assembly / negative regulation of viral genome replication / type I interferon-mediated signaling pathway / cellular response to exogenous dsRNA / cytoplasmic pattern recognition receptor signaling pathway / positive regulation of interferon-alpha production / antiviral innate immune response / TRAF6 mediated NF-kB activation / positive regulation of type I interferon production /  ubiquitin ligase complex / signaling adaptor activity / positive regulation of defense response to virus by host / activation of innate immune response / positive regulation of interferon-beta production / molecular condensate scaffold activity / Negative regulators of DDX58/IFIH1 signaling / positive regulation of interleukin-8 production / ubiquitin ligase complex / signaling adaptor activity / positive regulation of defense response to virus by host / activation of innate immune response / positive regulation of interferon-beta production / molecular condensate scaffold activity / Negative regulators of DDX58/IFIH1 signaling / positive regulation of interleukin-8 production /  mitochondrial membrane / DDX58/IFIH1-mediated induction of interferon-alpha/beta / PKR-mediated signaling / positive regulation of protein import into nucleus / positive regulation of interleukin-6 production / SARS-CoV-1 activates/modulates innate immune responses / positive regulation of DNA-binding transcription factor activity / Ovarian tumor domain proteases / positive regulation of tumor necrosis factor production / TRAF3-dependent IRF activation pathway / defense response to virus / positive regulation of canonical NF-kappaB signal transduction / mitochondrial outer membrane / molecular adaptor activity / defense response to bacterium / positive regulation of protein phosphorylation / mitochondrial membrane / DDX58/IFIH1-mediated induction of interferon-alpha/beta / PKR-mediated signaling / positive regulation of protein import into nucleus / positive regulation of interleukin-6 production / SARS-CoV-1 activates/modulates innate immune responses / positive regulation of DNA-binding transcription factor activity / Ovarian tumor domain proteases / positive regulation of tumor necrosis factor production / TRAF3-dependent IRF activation pathway / defense response to virus / positive regulation of canonical NF-kappaB signal transduction / mitochondrial outer membrane / molecular adaptor activity / defense response to bacterium / positive regulation of protein phosphorylation /  innate immune response / innate immune response /  protein kinase binding / SARS-CoV-2 activates/modulates innate and adaptive immune responses / protein kinase binding / SARS-CoV-2 activates/modulates innate and adaptive immune responses /  signal transduction / positive regulation of transcription by RNA polymerase II / signal transduction / positive regulation of transcription by RNA polymerase II /  mitochondrion / identical protein binding mitochondrion / identical protein bindingSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | ||||||
| Model details | lowest energy, model1 | ||||||
 Authors Authors | Spehr, J. / He, L. / Luehrs, T. / Ritter, C. | ||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 2016 Journal: Proc.Natl.Acad.Sci.USA / Year: 2016Title: Structure determination of helical filaments by solid-state NMR spectroscopy. Authors: He, L. / Bardiaux, B. / Ahmed, M. / Spehr, J. / Konig, R. / Lunsdorf, H. / Rand, U. / Luhrs, T. / Ritter, C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2ms8.cif.gz 2ms8.cif.gz | 633.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2ms8.ent.gz pdb2ms8.ent.gz | 529 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2ms8.json.gz 2ms8.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ms/2ms8 https://data.pdbj.org/pub/pdb/validation_reports/ms/2ms8 ftp://data.pdbj.org/pub/pdb/validation_reports/ms/2ms8 ftp://data.pdbj.org/pub/pdb/validation_reports/ms/2ms8 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 11817.363 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: MAVS, IPS1, KIAA1271, VISA / Production host: Homo sapiens (human) / Gene: MAVS, IPS1, KIAA1271, VISA / Production host:   Escherichia coli (E. coli) / References: UniProt: Q7Z434 Escherichia coli (E. coli) / References: UniProt: Q7Z434 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Contents: 300 uM [U-99% 13C; U-99% 15N] MAVS CARD-1, 93% H2O/7% D2O Solvent system: 93% H2O/7% D2O |
|---|---|
| Sample | Conc.: 300 uM / Component: MAVS CARD-1 / Isotopic labeling: [U-99% 13C; U-99% 15N] |
| Sample conditions | Ionic strength: 50 / pH: 3 / Pressure: ambient / Temperature: 298 K |
-NMR measurement
| NMR spectrometer | Type: Bruker Avance / Manufacturer: Bruker / Model : AVANCE / Field strength: 600 MHz : AVANCE / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 1 simulated annealing / Software ordinal: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||
| NMR constraints | NOE constraints total: 1098 | ||||||||||||||||||||||||||||||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: target function / Conformers calculated total number: 100 / Conformers submitted total number: 20 / Representative conformer: 1 |
 Movie
Movie Controller
Controller












 PDBj
PDBj








