+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2l90 | ||||||
|---|---|---|---|---|---|---|---|
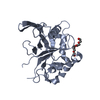
| Title | Solution structure of murine myristoylated msrA | ||||||
 Components Components | Peptide methionine sulfoxide reductase | ||||||
 Keywords Keywords |  OXIDOREDUCTASE OXIDOREDUCTASE | ||||||
| Function / homology |  Function and homology information Function and homology informationProtein repair / L-methionine-(S)-S-oxide reductase activity /  peptide-methionine (S)-S-oxide reductase / peptide-methionine (S)-S-oxide reductase /  peptide-methionine (S)-S-oxide reductase activity / peptide-methionine (S)-S-oxide reductase activity /  actin cytoskeleton / cellular response to oxidative stress / midbody / intracellular membrane-bounded organelle / actin cytoskeleton / cellular response to oxidative stress / midbody / intracellular membrane-bounded organelle /  mitochondrion / mitochondrion /  nucleoplasm ...Protein repair / L-methionine-(S)-S-oxide reductase activity / nucleoplasm ...Protein repair / L-methionine-(S)-S-oxide reductase activity /  peptide-methionine (S)-S-oxide reductase / peptide-methionine (S)-S-oxide reductase /  peptide-methionine (S)-S-oxide reductase activity / peptide-methionine (S)-S-oxide reductase activity /  actin cytoskeleton / cellular response to oxidative stress / midbody / intracellular membrane-bounded organelle / actin cytoskeleton / cellular response to oxidative stress / midbody / intracellular membrane-bounded organelle /  mitochondrion / mitochondrion /  nucleoplasm / nucleoplasm /  plasma membrane / plasma membrane /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Mus musculus (house mouse) Mus musculus (house mouse) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | ||||||
| Model details | minimized average, model 1 | ||||||
| Model type details | minimized average | ||||||
 Authors Authors | Gruschus, J.M. / Lim, J. / Piszczek, G. / Levine, R.L. / Tjandra, N. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2012 Journal: J.Biol.Chem. / Year: 2012Title: Characterization and solution structure of mouse myristoylated methionine sulfoxide reductase A. Authors: Lim, J.C. / Gruschus, J.M. / Ghesquiere, B. / Kim, G. / Piszczek, G. / Tjandra, N. / Levine, R.L. #1: Journal: J.Biol.Chem. / Year: 2012 Title: A Low pKa cysteine at the active site of mouse methionine sulfoxide reductase A. Authors: Lim, J.C. / Gruschus, J.M. / Kim, G. / Berlett, B.S. / Tjandra, N. / Levine, R.L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2l90.cif.gz 2l90.cif.gz | 1.3 MB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2l90.ent.gz pdb2l90.ent.gz | 1.1 MB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2l90.json.gz 2l90.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/l9/2l90 https://data.pdbj.org/pub/pdb/validation_reports/l9/2l90 ftp://data.pdbj.org/pub/pdb/validation_reports/l9/2l90 ftp://data.pdbj.org/pub/pdb/validation_reports/l9/2l90 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data | |
|---|---|
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 23679.570 Da / Num. of mol.: 1 / Fragment: sequence database residues 22-233 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Mus musculus (house mouse) / Gene: Msra / Production host: Mus musculus (house mouse) / Gene: Msra / Production host:   Escherichia coli (E. coli) Escherichia coli (E. coli)References: UniProt: Q9D6Y7,  peptide-methionine (S)-S-oxide reductase peptide-methionine (S)-S-oxide reductase |
|---|---|
| #2: Chemical | ChemComp-MYR /  Myristic acid Myristic acid |
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Contents: 0.3 mM [U-98% 13C; U-98% 15N] msrA, 5 mM DTT, 50 mM TRIS, 5 mM EDTA, 90% H2O/10% D2O Solvent system: 90% H2O/10% D2O | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||
| Sample conditions | pH: 7.0 / Pressure: ambient / Temperature: 300 K |
-NMR measurement
| NMR spectrometer | Type: Bruker Avance / Manufacturer: Bruker / Model : Avance / Field strength: 800 MHz : Avance / Field strength: 800 MHz |
|---|
- Processing
Processing
| NMR software |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 1 simulated annealing / Software ordinal: 1 Details: The authors state that the distortion around Cys206 CA in model 1 is a manifestation of the averaging, reflecting that local flexibility often cannot be accurately represented as a single structure. | |||||||||
| NMR representative | Selection criteria: minimized average structure | |||||||||
| NMR ensemble | Conformers calculated total number: 200 / Conformers submitted total number: 21 |
 Movie
Movie Controller
Controller











 PDBj
PDBj





