+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2e6m | ||||||
|---|---|---|---|---|---|---|---|
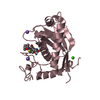
| Title | structure of mouse werner exonuclease domain | ||||||
 Components Components | Werner syndrome ATP-dependent helicase homolog | ||||||
 Keywords Keywords |  HYDROLASE / apo form HYDROLASE / apo form | ||||||
| Function / homology |  Function and homology information Function and homology informationProcessive synthesis on the C-strand of the telomere / Removal of the Flap Intermediate from the C-strand / Homologous DNA Pairing and Strand Exchange / Presynaptic phase of homologous DNA pairing and strand exchange / Resolution of D-loop Structures through Holliday Junction Intermediates / 3'-flap-structured DNA binding / positive regulation of strand invasion / HDR through Single Strand Annealing (SSA) / telomeric G-quadruplex DNA binding / HDR through Homologous Recombination (HRR) ...Processive synthesis on the C-strand of the telomere / Removal of the Flap Intermediate from the C-strand / Homologous DNA Pairing and Strand Exchange / Presynaptic phase of homologous DNA pairing and strand exchange / Resolution of D-loop Structures through Holliday Junction Intermediates / 3'-flap-structured DNA binding / positive regulation of strand invasion / HDR through Single Strand Annealing (SSA) / telomeric G-quadruplex DNA binding / HDR through Homologous Recombination (HRR) / forked DNA-dependent helicase activity / G2/M DNA damage checkpoint / 8-hydroxy-2'-deoxyguanosine DNA binding / telomeric D-loop binding / Processing of DNA double-strand break ends / regulation of growth rate / G-quadruplex DNA unwinding / telomeric D-loop disassembly / Y-form DNA binding / DNA 3'-5' helicase / four-way junction helicase activity / MutLalpha complex binding / Regulation of TP53 Activity through Phosphorylation / bubble DNA binding / protein localization to nucleolus / response to UV-C /  exonuclease activity / DNA metabolic process / 3'-5' DNA helicase activity / DNA synthesis involved in DNA repair / replication fork processing / DNA unwinding involved in DNA replication / exonuclease activity / DNA metabolic process / 3'-5' DNA helicase activity / DNA synthesis involved in DNA repair / replication fork processing / DNA unwinding involved in DNA replication /  replicative senescence / four-way junction DNA binding / replicative senescence / four-way junction DNA binding /  isomerase activity / cellular response to starvation / 3'-5' exonuclease activity / isomerase activity / cellular response to starvation / 3'-5' exonuclease activity /  DNA helicase activity / DNA helicase activity /  telomere maintenance / telomere maintenance /  replication fork / determination of adult lifespan / double-strand break repair via homologous recombination / replication fork / determination of adult lifespan / double-strand break repair via homologous recombination /  base-excision repair / cellular response to gamma radiation / base-excision repair / cellular response to gamma radiation /  cellular senescence / double-strand break repair / cellular senescence / double-strand break repair /  chromosome / manganese ion binding / regulation of apoptotic process / response to oxidative stress / chromosome / manganese ion binding / regulation of apoptotic process / response to oxidative stress /  DNA replication / DNA replication /  chromosome, telomeric region / chromosome, telomeric region /  Hydrolases; Acting on ester bonds / nuclear speck / Hydrolases; Acting on ester bonds / nuclear speck /  centrosome / centrosome /  chromatin binding / chromatin binding /  nucleolus / magnesium ion binding / protein homodimerization activity / nucleolus / magnesium ion binding / protein homodimerization activity /  ATP hydrolysis activity / ATP hydrolysis activity /  nucleoplasm / nucleoplasm /  ATP binding / ATP binding /  nucleus / nucleus /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Mus musculus (house mouse) Mus musculus (house mouse) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | ||||||
 Authors Authors | Cho, Y. / Choi, J.M. | ||||||
 Citation Citation |  Journal: TO BE PUBLISHED Journal: TO BE PUBLISHEDTitle: probing the roles of active site residues in 3'-5' exonuclease of werner syndrome protein Authors: Cho, Y. / Choi, J.M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2e6m.cif.gz 2e6m.cif.gz | 52.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2e6m.ent.gz pdb2e6m.ent.gz | 36.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2e6m.json.gz 2e6m.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/e6/2e6m https://data.pdbj.org/pub/pdb/validation_reports/e6/2e6m ftp://data.pdbj.org/pub/pdb/validation_reports/e6/2e6m ftp://data.pdbj.org/pub/pdb/validation_reports/e6/2e6m | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2e6lSC S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 23401.854 Da / Num. of mol.: 1 / Fragment: wrn exonuclease domain, residues 31-238 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Mus musculus (house mouse) / Gene: Wrn / Plasmid: pET-28a / Species (production host): Escherichia coli / Production host: Mus musculus (house mouse) / Gene: Wrn / Plasmid: pET-28a / Species (production host): Escherichia coli / Production host:   Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21(DE3) Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21(DE3)References: UniProt: O09053,  Hydrolases; Acting on acid anhydrides; In phosphorus-containing anhydrides Hydrolases; Acting on acid anhydrides; In phosphorus-containing anhydrides |
|---|---|
| #2: Chemical | ChemComp-SO4 /  Sulfate Sulfate |
| #3: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.1 Å3/Da / Density % sol: 41.41 % |
|---|---|
Crystal grow | Temperature: 291.15 K / Method: vapor diffusion, hanging drop / pH: 8.5 Details: pH 8.5, VAPOR DIFFUSION, HANGING DROP, temperature 291.15K |
-Data collection
| Diffraction | Mean temperature: 93.15 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: PAL/PLS SYNCHROTRON / Site: PAL/PLS  / Beamline: 4A / Wavelength: 1 Å / Beamline: 4A / Wavelength: 1 Å |
| Detector | Detector: CCD / Date: Feb 14, 2005 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1 Å / Relative weight: 1 : 1 Å / Relative weight: 1 |
| Reflection | Resolution: 2→50 Å / Num. all: 14814 / Num. obs: 14814 / % possible obs: 96 % / Redundancy: 5.2 % / Biso Wilson estimate: 7 Å2 / Rsym value: 0.092 / Net I/σ(I): 29 |
| Reflection shell | Resolution: 2→2.11 Å / Redundancy: 4.7 % / Mean I/σ(I) obs: 29 / Num. unique all: 14814 / Rsym value: 0.092 / % possible all: 96 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 2E6L Resolution: 2→29.6 Å / Rfactor Rfree error: 0.01 / Data cutoff high absF: 77755.73 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 1
| ||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 51.3719 Å2 / ksol: 0.390643 e/Å3 | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 25 Å2
| ||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→29.6 Å
| ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2→2.13 Å / Rfactor Rfree error: 0.028 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj






