[English] 日本語
 Yorodumi
Yorodumi- PDB-2dp7: Crystal Structure of D(CGCGAATXCGCG) Where X is 5-(N-aminohexyl)c... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2dp7 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Crystal Structure of D(CGCGAATXCGCG) Where X is 5-(N-aminohexyl)carbamoyl-2'-deoxyuridine | ||||||||||||||||||
 Components Components | DNA (5'-D(* Keywords Keywords DNA / modified nucleotide / 5-(N-aminohexyl)carbamoyl-2'-deoxyuridine / DNA / modified nucleotide / 5-(N-aminohexyl)carbamoyl-2'-deoxyuridine /  B-DNA B-DNAFunction / homology | (6-AMINOHEXYL)CARBAMIC ACID / : / |  DNA / DNA (> 10) DNA / DNA (> 10) Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.55 Å MOLECULAR REPLACEMENT / Resolution: 1.55 Å  Authors AuthorsJuan, E.C.M. / Kondo, J. / Kurihara, T. / Ito, T. / Ueno, Y. / Matsuda, A. / Takenaka, A. |  Citation Citation Journal: Nucleic Acids Res. / Year: 2007 Journal: Nucleic Acids Res. / Year: 2007Title: Crystal structures of DNA:DNA and DNA:RNA duplexes containing 5-(N-aminohexyl)carbamoyl-modified uracils reveal the basis for properties as antigene and antisense molecules Authors: Juan, E.C.M. / Kondo, J. / Kurihara, T. / Ito, T. / Ueno, Y. / Matsuda, A. / Takenaka, A. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2dp7.cif.gz 2dp7.cif.gz | 29.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2dp7.ent.gz pdb2dp7.ent.gz | 19.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2dp7.json.gz 2dp7.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dp/2dp7 https://data.pdbj.org/pub/pdb/validation_reports/dp/2dp7 ftp://data.pdbj.org/pub/pdb/validation_reports/dp/2dp7 ftp://data.pdbj.org/pub/pdb/validation_reports/dp/2dp7 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2dpbC  2dpcC  2dqoC  2dqpC  2dqqC  355dS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 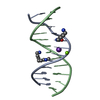
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 3649.366 Da / Num. of mol.: 2 / Source method: obtained synthetically / Details: This DNA dodecamer is a synthetic construct. #2: Chemical | ChemComp-MG / | #3: Chemical | #4: Chemical | ChemComp-K / | #5: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.16 Å3/Da / Density % sol: 42.97 % | ||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 6 Details: 20mM sodium cacodylate (pH 6.0), 40mM potassium chloride, 2.5mM magnesium chloride, 1.5mM spermine tetrahydrochloride and 5%(v/v) 2-methyl-2,4-pentanediol (MPD), equilibrated against 45%(v/v) ...Details: 20mM sodium cacodylate (pH 6.0), 40mM potassium chloride, 2.5mM magnesium chloride, 1.5mM spermine tetrahydrochloride and 5%(v/v) 2-methyl-2,4-pentanediol (MPD), equilibrated against 45%(v/v) MPD, VAPOR DIFFUSION, HANGING DROP, temperature 277K | ||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions |
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Photon Factory Photon Factory  / Beamline: BL-18B / Wavelength: 1 Å / Beamline: BL-18B / Wavelength: 1 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Jan 18, 2002 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1 Å / Relative weight: 1 : 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.55→18.836 Å / Num. obs: 10074 / % possible obs: 100 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Rmerge(I) obs: 0.047 / Net I/σ(I): 8.4 |
| Reflection shell | Resolution: 1.55→1.63 Å / Rmerge(I) obs: 0.257 / Mean I/σ(I) obs: 2.8 / % possible all: 100 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 355D Resolution: 1.55→10 Å / σ(F): 3 / Stereochemistry target values: Maximum Likelihood Details: The value of the item "Number reflections (observed)" is the number of reflections used in refinement. This value excludes the reflections that have sigma(F) less than 3.0.
| |||||||||||||||||||||||||
| Displacement parameters |
| |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.55→10 Å
|
 Movie
Movie Controller
Controller




 PDBj
PDBj