[English] 日本語
 Yorodumi
Yorodumi- PDB-2b19: Solution Structure of mammalian tachykinin peptide, Neuropeptide K -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2b19 | ||||||
|---|---|---|---|---|---|---|---|
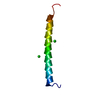
| Title | Solution Structure of mammalian tachykinin peptide, Neuropeptide K | ||||||
 Components Components | Neuropeptide K | ||||||
 Keywords Keywords |  NEUROPEPTIDE / NEUROPEPTIDE /  helix / helix /  3 10 helix / lipid induced conformation / DPC micelles 3 10 helix / lipid induced conformation / DPC micelles | ||||||
| Function / homology |  Function and homology information Function and homology informationneuromedin K receptor binding / substance K receptor binding / positive regulation of sensory perception of pain / positive regulation of corticosterone secretion / substance P receptor binding /  insemination / insemination /  : / : /  : / positive regulation of flagellated sperm motility / positive regulation of saliva secretion ...neuromedin K receptor binding / substance K receptor binding / positive regulation of sensory perception of pain / positive regulation of corticosterone secretion / substance P receptor binding / : / positive regulation of flagellated sperm motility / positive regulation of saliva secretion ...neuromedin K receptor binding / substance K receptor binding / positive regulation of sensory perception of pain / positive regulation of corticosterone secretion / substance P receptor binding /  insemination / insemination /  : / : /  : / positive regulation of flagellated sperm motility / positive regulation of saliva secretion / positive regulation of renal sodium excretion / Tachykinin receptors bind tachykinins / detection of abiotic stimulus / positive regulation of synaptic transmission, cholinergic / positive regulation of lymphocyte proliferation / tachykinin receptor signaling pathway / response to yeast / positive regulation of action potential / regulation of sensory perception of pain / positive regulation of acute inflammatory response / antifungal humoral response / response to morphine / positive regulation of ossification / negative regulation of systemic arterial blood pressure / negative regulation of heart rate / response to pain / positive regulation of epithelial cell migration / detection of temperature stimulus involved in sensory perception of pain / : / positive regulation of flagellated sperm motility / positive regulation of saliva secretion / positive regulation of renal sodium excretion / Tachykinin receptors bind tachykinins / detection of abiotic stimulus / positive regulation of synaptic transmission, cholinergic / positive regulation of lymphocyte proliferation / tachykinin receptor signaling pathway / response to yeast / positive regulation of action potential / regulation of sensory perception of pain / positive regulation of acute inflammatory response / antifungal humoral response / response to morphine / positive regulation of ossification / negative regulation of systemic arterial blood pressure / negative regulation of heart rate / response to pain / positive regulation of epithelial cell migration / detection of temperature stimulus involved in sensory perception of pain /  associative learning / neuropeptide signaling pathway / associative learning / neuropeptide signaling pathway /  long-term memory / positive regulation of stress fiber assembly / sensory perception of pain / response to hormone / cellular response to nerve growth factor stimulus / positive regulation of synaptic transmission, GABAergic / long-term memory / positive regulation of stress fiber assembly / sensory perception of pain / response to hormone / cellular response to nerve growth factor stimulus / positive regulation of synaptic transmission, GABAergic /  regulation of blood pressure / antimicrobial humoral immune response mediated by antimicrobial peptide / cell-cell signaling / antibacterial humoral response / positive regulation of cytosolic calcium ion concentration / chemical synaptic transmission / G alpha (q) signalling events / killing of cells of another organism / defense response to Gram-negative bacterium / response to lipopolysaccharide / regulation of blood pressure / antimicrobial humoral immune response mediated by antimicrobial peptide / cell-cell signaling / antibacterial humoral response / positive regulation of cytosolic calcium ion concentration / chemical synaptic transmission / G alpha (q) signalling events / killing of cells of another organism / defense response to Gram-negative bacterium / response to lipopolysaccharide /  receptor ligand activity / defense response to Gram-positive bacterium / receptor ligand activity / defense response to Gram-positive bacterium /  inflammatory response / G protein-coupled receptor signaling pathway / inflammatory response / G protein-coupled receptor signaling pathway /  axon / axon /  innate immune response / neuronal cell body / innate immune response / neuronal cell body /  synapse / synapse /  extracellular space / extracellular region / extracellular space / extracellular region /  plasma membrane plasma membraneSimilarity search - Function | ||||||
| Method |  SOLUTION NMR / distance geometry, simulated annealing (DYANA) SOLUTION NMR / distance geometry, simulated annealing (DYANA) | ||||||
 Authors Authors | Dike, A. / Cowsik, S.M. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2006 Journal: Biochemistry / Year: 2006Title: Three-dimensional structure of neuropeptide k bound to dodecylphosphocholine micelles Authors: Dike, A. / Cowsik, S.M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2b19.cif.gz 2b19.cif.gz | 222.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2b19.ent.gz pdb2b19.ent.gz | 185.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2b19.json.gz 2b19.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/b1/2b19 https://data.pdbj.org/pub/pdb/validation_reports/b1/2b19 ftp://data.pdbj.org/pub/pdb/validation_reports/b1/2b19 ftp://data.pdbj.org/pub/pdb/validation_reports/b1/2b19 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 3989.559 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: Chemical Synthesis / References: UniProt: Q53GH4, UniProt: P20366*PLUS |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||
| NMR details | Text: This structure was determined using standard 2D homonuclear techniques |
- Sample preparation
Sample preparation
| Details | Contents: 146mM perdeuterated DPC; 90% H2O, 10% D2O / Solvent system: 90% H2O/10% D2O |
|---|---|
| Sample conditions | Ionic strength: No salts used / pH: 3.5 / Pressure: ambient / Temperature: 305 K |
-NMR measurement
| NMR spectrometer | Type: Bruker DRX / Manufacturer: Bruker / Model : DRX / Field strength: 500 MHz : DRX / Field strength: 500 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: distance geometry, simulated annealing (DYANA) / Software ordinal: 1 | ||||||||||||||||
| NMR representative | Selection criteria: fewest violations | ||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the least restraint violations Conformers calculated total number: 50 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller










 PDBj
PDBj