+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1wou | ||||||
|---|---|---|---|---|---|---|---|
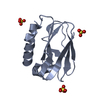
| Title | Crystal Structure of human Trp14 | ||||||
 Components Components | thioredoxin -related protein, 14 kDa | ||||||
 Keywords Keywords |  ELECTRON TRANSPORT ELECTRON TRANSPORT | ||||||
| Function / homology |  Function and homology information Function and homology informationprotein-disulfide reductase (NAD(P)H) activity / tumor necrosis factor-mediated signaling pathway /  peroxidase activity / extracellular exosome / peroxidase activity / extracellular exosome /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MAD / Resolution: 1.8 Å MAD / Resolution: 1.8 Å | ||||||
 Authors Authors | Woo, J.R. / Kim, S.J. / Jeong, W. / Cho, Y.H. / Lee, S.C. / Chung, Y.J. / Rhee, S.G. / Ryu, S.E. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2004 Journal: J.Biol.Chem. / Year: 2004Title: Structural basis of cellular redox regulation by human TRP14 Authors: Woo, J.R. / Kim, S.J. / Jeong, W. / Cho, Y.H. / Lee, S.C. / Chung, Y.J. / Rhee, S.G. / Ryu, S.E. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1wou.cif.gz 1wou.cif.gz | 29.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1wou.ent.gz pdb1wou.ent.gz | 23.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1wou.json.gz 1wou.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/wo/1wou https://data.pdbj.org/pub/pdb/validation_reports/wo/1wou ftp://data.pdbj.org/pub/pdb/validation_reports/wo/1wou ftp://data.pdbj.org/pub/pdb/validation_reports/wo/1wou | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 13957.786 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Plasmid: pET11a / Production host: Homo sapiens (human) / Plasmid: pET11a / Production host:   Escherichia coli (E. coli) / References: UniProt: Q9BRA2 Escherichia coli (E. coli) / References: UniProt: Q9BRA2 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 2 X-RAY DIFFRACTION / Number of used crystals: 2 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.97 Å3/Da / Density % sol: 37 % |
|---|---|
Crystal grow | Temperature: 291 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: polyethylene glycol 8000, polyethylene glycol monomethylether 2000, pH 6.5, VAPOR DIFFUSION, HANGING DROP, temperature 291K |
-Data collection
| Diffraction |
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RU300 / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU RU300 / Wavelength: 1.5418 Å | ||||||||||||||||||
| Detector |
| ||||||||||||||||||
| Radiation |
| ||||||||||||||||||
| Radiation wavelength | Wavelength : 1.5418 Å / Relative weight: 1 : 1.5418 Å / Relative weight: 1 | ||||||||||||||||||
| Reflection | Resolution: 1.8→40 Å / Num. all: 10425 / Num. obs: 10385 / % possible obs: 99.6 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Rmerge(I) obs: 0.041 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MAD / Resolution: 1.8→22.57 Å / Isotropic thermal model: isothermal / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / Stereochemistry target values: Engh & Huber MAD / Resolution: 1.8→22.57 Å / Isotropic thermal model: isothermal / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / Stereochemistry target values: Engh & Huber
| |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→22.57 Å
|
 Movie
Movie Controller
Controller












 PDBj
PDBj