+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ukr | ||||||
|---|---|---|---|---|---|---|---|
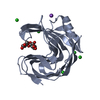
| Title | STRUCTURE OF ENDO-1,4-BETA-XYLANASE C | ||||||
 Components Components | ENDO-1,4-B-XYLANASE I | ||||||
 Keywords Keywords |  HYDROLASE / XYLAN DEGRADATION / GLYCOSIDASE HYDROLASE / XYLAN DEGRADATION / GLYCOSIDASE | ||||||
| Function / homology |  Function and homology information Function and homology information endo-1,4-beta-xylanase activity / endo-1,4-beta-xylanase activity /  endo-1,4-beta-xylanase / xylan catabolic process / extracellular region endo-1,4-beta-xylanase / xylan catabolic process / extracellular regionSimilarity search - Function | ||||||
| Biological species |   Aspergillus niger (mold) Aspergillus niger (mold) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.4 Å MOLECULAR REPLACEMENT / Resolution: 2.4 Å | ||||||
 Authors Authors | Krengel, U. / Dijkstra, B.W. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1996 Journal: J.Mol.Biol. / Year: 1996Title: Three-dimensional structure of Endo-1,4-beta-xylanase I from Aspergillus niger: molecular basis for its low pH optimum. Authors: Krengel, U. / Dijkstra, B.W. #1:  Journal: Acta Crystallogr.,Sect.D / Year: 1996 Journal: Acta Crystallogr.,Sect.D / Year: 1996Title: Crystallization and Preliminary Crystallographic Analysis of Endo-1,4-Beta-Xyalanase I from Aspergillus Niger Authors: Krengel, U. / Rozeboom, H.J. / Kalk, K.H. / Dijkstra, B.W. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ukr.cif.gz 1ukr.cif.gz | 145.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ukr.ent.gz pdb1ukr.ent.gz | 120.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ukr.json.gz 1ukr.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/uk/1ukr https://data.pdbj.org/pub/pdb/validation_reports/uk/1ukr ftp://data.pdbj.org/pub/pdb/validation_reports/uk/1ukr ftp://data.pdbj.org/pub/pdb/validation_reports/uk/1ukr | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 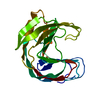 1xndS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| 4 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 19855.941 Da / Num. of mol.: 4 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Aspergillus niger (mold) / Production host: Aspergillus niger (mold) / Production host:   Escherichia coli (E. coli) / References: UniProt: P55329, Escherichia coli (E. coli) / References: UniProt: P55329,  endo-1,4-beta-xylanase endo-1,4-beta-xylanase#2: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.57 Å3/Da / Density % sol: 52.1 % | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Method: vapor diffusion - sitting drop - macroseeding / pH: 4.7 Details: SITTING DROP SETUP USING SILICONIZED GLASS WELLS USING 15 MICROLITER DROPS AT ROOM TEMPERATURE AND MACROSEEDING. PROTEIN SOLUTION: 4.5 MG/ML IN 10 MM NA ACETATE PH 4.7. RESERVOIR SOLUTION: 1. ...Details: SITTING DROP SETUP USING SILICONIZED GLASS WELLS USING 15 MICROLITER DROPS AT ROOM TEMPERATURE AND MACROSEEDING. PROTEIN SOLUTION: 4.5 MG/ML IN 10 MM NA ACETATE PH 4.7. RESERVOIR SOLUTION: 1.5 ML 1.8 M NA2S2O3 (PH 8.0)., vapor diffusion - sitting drop - macroseeding Temp details: room temp | |||||||||||||||
| Crystal grow | *PLUS Temperature: 295, 277, 310 K / Method: vapor diffusion, hanging drop / Details: or sitting drop vapor diffusion or batch method | |||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 295 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  EMBL/DESY, HAMBURG EMBL/DESY, HAMBURG  / Beamline: X31 / Wavelength: 0.99 / Beamline: X31 / Wavelength: 0.99 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Oct 20, 1994 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.99 Å / Relative weight: 1 : 0.99 Å / Relative weight: 1 |
| Reflection | Resolution: 2.35→38.06 Å / Num. obs: 33131 / % possible obs: 95.3 % / Redundancy: 4.6 % / Rsym value: 0.082 |
| Reflection | *PLUS Rmerge(I) obs: 0.082 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: TRICHODERMA HARZIANUM XYLANASE, PDB ENTRY 1XND. Resolution: 2.4→6 Å / σ(F): 0 Details: REFINEMENT INCLUDED NCS-RESTRAINTS AT PLACES WITHOUT CRYSTAL CONTACTS. ONLY IN THE FINAL REFINEMENT CYCLE ALL DATA WERE INCLUDED. IN THE EARLIER STAGES OF REFINEMENT, 10% OF THE DATA WERE ...Details: REFINEMENT INCLUDED NCS-RESTRAINTS AT PLACES WITHOUT CRYSTAL CONTACTS. ONLY IN THE FINAL REFINEMENT CYCLE ALL DATA WERE INCLUDED. IN THE EARLIER STAGES OF REFINEMENT, 10% OF THE DATA WERE TAKEN TO CALCULATE A FREE R-FACTOR (R=17.7 AND RFREE=21.9 %, RESPECTIVELY).
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 19.2 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.24 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.4→6 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.1 / Classification: refinement X-PLOR / Version: 3.1 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller












 PDBj
PDBj

