+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1tk7 | ||||||
|---|---|---|---|---|---|---|---|
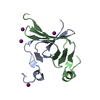
| Title | NMR structure of WW domains (WW3-4) from Suppressor of Deltex | ||||||
 Components Components | CG4244-PB | ||||||
 Keywords Keywords |  SIGNALING PROTEIN / SIGNALING PROTEIN /  WW domain / Notch WW domain / Notch | ||||||
| Function / homology |  Function and homology information Function and homology informationpupal development / RHOQ GTPase cycle / imaginal disc-derived leg joint morphogenesis / R8 cell development / imaginal disc-derived wing margin morphogenesis / Activated NOTCH1 Transmits Signal to the Nucleus / RUNX1 regulates transcription of genes involved in differentiation of HSCs / wing disc dorsal/ventral pattern formation / Downregulation of ERBB4 signaling / RHOU GTPase cycle ...pupal development / RHOQ GTPase cycle / imaginal disc-derived leg joint morphogenesis / R8 cell development / imaginal disc-derived wing margin morphogenesis / Activated NOTCH1 Transmits Signal to the Nucleus / RUNX1 regulates transcription of genes involved in differentiation of HSCs / wing disc dorsal/ventral pattern formation / Downregulation of ERBB4 signaling / RHOU GTPase cycle / imaginal disc-derived wing vein morphogenesis / Regulation of PTEN stability and activity / Antigen processing: Ubiquitination & Proteasome degradation / protein targeting to lysosome /  lipid storage / HECT-type E3 ubiquitin transferase / Notch binding / negative regulation of hippo signaling / negative regulation of Notch signaling pathway / positive regulation of endocytosis / negative regulation of smoothened signaling pathway / lipid storage / HECT-type E3 ubiquitin transferase / Notch binding / negative regulation of hippo signaling / negative regulation of Notch signaling pathway / positive regulation of endocytosis / negative regulation of smoothened signaling pathway /  Notch signaling pathway / Notch signaling pathway /  receptor internalization / ubiquitin-protein transferase activity / receptor internalization / ubiquitin-protein transferase activity /  regulation of protein localization / regulation of protein localization /  ubiquitin protein ligase activity / ubiquitin protein ligase activity /  cell cortex / proteasome-mediated ubiquitin-dependent protein catabolic process / protein ubiquitination / cell cortex / proteasome-mediated ubiquitin-dependent protein catabolic process / protein ubiquitination /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Drosophila melanogaster (fruit fly) Drosophila melanogaster (fruit fly) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing, torsion angle dynamics simulated annealing, torsion angle dynamics | ||||||
 Authors Authors | Fedoroff, O.Y. / Avis, J.M. / Golovanov, A.P. / Baron, M. / Townson, S.A. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2004 Journal: J.Biol.Chem. / Year: 2004Title: The structure and dynamics of tandem WW domains in a negative regulator of notch signaling, Suppressor of deltex Authors: Fedoroff, O.Y. / Townson, S.A. / Golovanov, A.P. / Baron, M. / Avis, J.M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1tk7.cif.gz 1tk7.cif.gz | 224.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1tk7.ent.gz pdb1tk7.ent.gz | 187.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1tk7.json.gz 1tk7.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/tk/1tk7 https://data.pdbj.org/pub/pdb/validation_reports/tk/1tk7 ftp://data.pdbj.org/pub/pdb/validation_reports/tk/1tk7 ftp://data.pdbj.org/pub/pdb/validation_reports/tk/1tk7 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 10280.271 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Drosophila melanogaster (fruit fly) / Gene: Su(dx) / Plasmid: pGEX / Production host: Drosophila melanogaster (fruit fly) / Gene: Su(dx) / Plasmid: pGEX / Production host:   Escherichia coli (E. coli) / Strain (production host): BCl21 / References: UniProt: Q9Y0H4 Escherichia coli (E. coli) / Strain (production host): BCl21 / References: UniProt: Q9Y0H4 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Contents: 0.5 PBS, 50mM Arg/Glu / Solvent system: 90% H2O/10% D2O |
|---|---|
| Sample conditions | Ionic strength: 150 Na/KCl / pH: 7.1 / Pressure: 1 atm / Temperature: 288 K |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| NMR spectrometer | Type: Bruker AMX / Manufacturer: Bruker / Model : AMX / Field strength: 600 MHz : AMX / Field strength: 600 MHz |
- Processing
Processing
| NMR software | Name: ARIA / Version: 1.2 / Developer: Nilges / Classification: refinement |
|---|---|
| Refinement | Method:  simulated annealing, torsion angle dynamics / Software ordinal: 1 simulated annealing, torsion angle dynamics / Software ordinal: 1 |
| NMR representative | Selection criteria: lowest energy |
| NMR ensemble | Conformer selection criteria: structures with favorable non-bond energy Conformers calculated total number: 20 / Conformers submitted total number: 7 |
 Movie
Movie Controller
Controller










 PDBj
PDBj



