[English] 日本語
 Yorodumi
Yorodumi- PDB-1rng: SOLUTION STRUCTURE OF THE CUUG HAIRPIN: A NOVEL RNA TETRALOOP MOTIF -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1rng | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
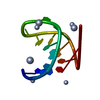
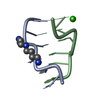
| Title | SOLUTION STRUCTURE OF THE CUUG HAIRPIN: A NOVEL RNA TETRALOOP MOTIF | ||||||||||||||||||
 Components Components | RNA (5'-R(* Keywords Keywords RNA / RNA /  TETRALOOP TETRALOOPFunction / homology |  RNA / RNA (> 10) RNA / RNA (> 10) Function and homology information Function and homology informationMethod |  SOLUTION NMR SOLUTION NMR  Authors AuthorsJucker, F.M. / Pardi, A. |  Citation Citation Journal: Biochemistry / Year: 1995 Journal: Biochemistry / Year: 1995Title: Solution structure of the CUUG hairpin loop: a novel RNA tetraloop motif. Authors: Jucker, F.M. / Pardi, A. #1:  Journal: Proc.Natl.Acad.Sci.USA / Year: 1990 Journal: Proc.Natl.Acad.Sci.USA / Year: 1990Title: Architecture of Ribosomal RNA: Constraints on the Sequence of "Tetra-Loops" Authors: Woese, C.R. / Winker, S. / Gutell, R.R. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1rng.cif.gz 1rng.cif.gz | 43.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1rng.ent.gz pdb1rng.ent.gz | 31.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1rng.json.gz 1rng.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/rn/1rng https://data.pdbj.org/pub/pdb/validation_reports/rn/1rng ftp://data.pdbj.org/pub/pdb/validation_reports/rn/1rng ftp://data.pdbj.org/pub/pdb/validation_reports/rn/1rng | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 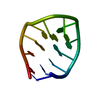
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: RNA chain | Mass: 3820.296 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: OLIGONUCLEOTIDE MADE BY IN VITRO TRANSCRIPTION /  Keywords: RIBONUCLEIC ACID Keywords: RIBONUCLEIC ACID |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR |
|---|
- Sample preparation
Sample preparation
Crystal grow | *PLUS Method: other |
|---|
- Processing
Processing
| Software |
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| NMR software | Name:  X-PLOR / Developer: BRUNGER / Classification: refinement X-PLOR / Developer: BRUNGER / Classification: refinement | ||||||||
| Refinement | Software ordinal: 1 / Details: NUCLEIC ACID PARAMETER SET. | ||||||||
| NMR ensemble | Conformers submitted total number: 5 |
 Movie
Movie Controller
Controller








 PDBj
PDBj





























