+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1q7d | ||||||
|---|---|---|---|---|---|---|---|
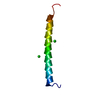
| Title | Structure of the integrin alpha2beta1 binding collagen peptide | ||||||
 Components Components | collagen alfa 1(I) chain peptide GPOGPOGFOGERGPOGPOGPO | ||||||
 Keywords Keywords |  CONTRACTILE PROTEIN / CONTRACTILE PROTEIN /  COLLAGEN / COLLAGEN /  INTEGRIN / TRIPLEHELIX INTEGRIN / TRIPLEHELIX | ||||||
| Function / homology |  Function and homology information Function and homology informationcellular response to fluoride / collagen type I trimer / tooth mineralization / cellular response to vitamin E / collagen type IV trimer / Anchoring fibril formation / Crosslinking of collagen fibrils / collagen biosynthetic process / Collagen chain trimerization / Enhanced cleavage of VWF variant by ADAMTS13 ...cellular response to fluoride / collagen type I trimer / tooth mineralization / cellular response to vitamin E / collagen type IV trimer / Anchoring fibril formation / Crosslinking of collagen fibrils / collagen biosynthetic process / Collagen chain trimerization / Enhanced cleavage of VWF variant by ADAMTS13 / Defective VWF cleavage by ADAMTS13 variant / Defective VWF binding to collagen type I /  platelet-derived growth factor binding / bone trabecula formation / extracellular matrix structural constituent conferring tensile strength / Enhanced binding of GP1BA variant to VWF multimer:collagen / Defective binding of VWF variant to GPIb:IX:V / platelet-derived growth factor binding / bone trabecula formation / extracellular matrix structural constituent conferring tensile strength / Enhanced binding of GP1BA variant to VWF multimer:collagen / Defective binding of VWF variant to GPIb:IX:V /  intramembranous ossification / Extracellular matrix organization / embryonic skeletal system development / Collagen biosynthesis and modifying enzymes / cartilage development involved in endochondral bone morphogenesis / skin morphogenesis / collagen-activated tyrosine kinase receptor signaling pathway / Platelet Adhesion to exposed collagen / intramembranous ossification / Extracellular matrix organization / embryonic skeletal system development / Collagen biosynthesis and modifying enzymes / cartilage development involved in endochondral bone morphogenesis / skin morphogenesis / collagen-activated tyrosine kinase receptor signaling pathway / Platelet Adhesion to exposed collagen /  endochondral ossification / cellular response to fibroblast growth factor stimulus / collagen fibril organization / negative regulation of cell-substrate adhesion / response to steroid hormone / face morphogenesis / Scavenging by Class A Receptors / skin development / MET activates PTK2 signaling / Assembly of collagen fibrils and other multimeric structures / Syndecan interactions / GP1b-IX-V activation signalling / blood vessel development / RUNX2 regulates osteoblast differentiation / Platelet Aggregation (Plug Formation) / Collagen degradation / protein localization to nucleus / Non-integrin membrane-ECM interactions / ECM proteoglycans / response to hyperoxia / Integrin cell surface interactions / positive regulation of epithelial to mesenchymal transition / response to mechanical stimulus / cellular response to retinoic acid / GPVI-mediated activation cascade / cellular response to epidermal growth factor stimulus / response to cAMP / cellular response to transforming growth factor beta stimulus / endochondral ossification / cellular response to fibroblast growth factor stimulus / collagen fibril organization / negative regulation of cell-substrate adhesion / response to steroid hormone / face morphogenesis / Scavenging by Class A Receptors / skin development / MET activates PTK2 signaling / Assembly of collagen fibrils and other multimeric structures / Syndecan interactions / GP1b-IX-V activation signalling / blood vessel development / RUNX2 regulates osteoblast differentiation / Platelet Aggregation (Plug Formation) / Collagen degradation / protein localization to nucleus / Non-integrin membrane-ECM interactions / ECM proteoglycans / response to hyperoxia / Integrin cell surface interactions / positive regulation of epithelial to mesenchymal transition / response to mechanical stimulus / cellular response to retinoic acid / GPVI-mediated activation cascade / cellular response to epidermal growth factor stimulus / response to cAMP / cellular response to transforming growth factor beta stimulus /  visual perception / extracellular matrix organization / visual perception / extracellular matrix organization /  ossification / ossification /  secretory granule / secretory granule /  skeletal system development / Cell surface interactions at the vascular wall / cellular response to glucose stimulus / sensory perception of sound / cellular response to amino acid stimulus / response to insulin / response to hydrogen peroxide / osteoblast differentiation / cellular response to mechanical stimulus / Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell / positive regulation of canonical Wnt signaling pathway / skeletal system development / Cell surface interactions at the vascular wall / cellular response to glucose stimulus / sensory perception of sound / cellular response to amino acid stimulus / response to insulin / response to hydrogen peroxide / osteoblast differentiation / cellular response to mechanical stimulus / Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell / positive regulation of canonical Wnt signaling pathway /  protein transport / response to estradiol / cellular response to tumor necrosis factor / collagen-containing extracellular matrix / protein transport / response to estradiol / cellular response to tumor necrosis factor / collagen-containing extracellular matrix /  protease binding / positive regulation of cell migration / response to xenobiotic stimulus / protease binding / positive regulation of cell migration / response to xenobiotic stimulus /  endoplasmic reticulum lumen / positive regulation of DNA-templated transcription / endoplasmic reticulum lumen / positive regulation of DNA-templated transcription /  extracellular space / extracellular region / identical protein binding / extracellular space / extracellular region / identical protein binding /  metal ion binding / metal ion binding /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å MOLECULAR REPLACEMENT / Resolution: 1.8 Å | ||||||
 Authors Authors | Emsley, J. / Knight, C.G. / Farndale, R.W. / Barnes, M.J. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2004 Journal: J.Mol.Biol. / Year: 2004Title: Structure of the Integrin alpha2beta1-binding Collagen Peptide. Authors: Emsley, J. / Knight, C.G. / Farndale, R.W. / Barnes, M.J. | ||||||
| History |
| ||||||
| Remark 999 | SEQUENCE The peptide is acetylated at the N-terminus and amidated at the C-terminus. Gly-Pro-Hyp ...SEQUENCE The peptide is acetylated at the N-terminus and amidated at the C-terminus. Gly-Pro-Hyp groups were added for stabilization. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1q7d.cif.gz 1q7d.cif.gz | 20.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1q7d.ent.gz pdb1q7d.ent.gz | 17.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1q7d.json.gz 1q7d.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/q7/1q7d https://data.pdbj.org/pub/pdb/validation_reports/q7/1q7d ftp://data.pdbj.org/pub/pdb/validation_reports/q7/1q7d ftp://data.pdbj.org/pub/pdb/validation_reports/q7/1q7d | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||||||||
| Unit cell |
| |||||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein/peptide | Mass: 2039.167 Da / Num. of mol.: 3 / Source method: obtained synthetically Details: THE PEPTIDE WAS CHEMICALLY SYNTHESIZED. THE SEQUENCE OF THE PEPTIDE IS NATURALLY FOUND IN HOMO SAPIENS (HUMAN). PEPTIDE SYNTHESISED USING STANDARD FMOC SYNTHESIS References: UniProt: P02452 #2: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.12 Å3/Da / Density % sol: 41.66 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Temperature: 277 K / Method: vapor diffusion, sitting drop / pH: 8 Details: PEG 8K, pH 8.0, VAPOR DIFFUSION, SITTING DROP, temperature 277K | ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / pH: 6.5 / Method: vapor diffusion, sitting drop | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 85 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID13 / Wavelength: 1 Å / Beamline: ID13 / Wavelength: 1 Å |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Jan 6, 2001 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1 Å / Relative weight: 1 : 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.8→20 Å / Num. all: 5060 / Num. obs: 5060 / % possible obs: 99.1 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 6.6 % / Biso Wilson estimate: 0.8 Å2 / Rmerge(I) obs: 0.105 / Rsym value: 0.105 / Net I/σ(I): 12.9 |
| Reflection shell | Resolution: 1.8→1.9 Å / Redundancy: 4.8 % / Rmerge(I) obs: 0.355 / Mean I/σ(I) obs: 5 / Num. unique all: 689 / Rsym value: 0.355 / % possible all: 95.2 |
| Reflection | *PLUS % possible obs: 98.2 % / Redundancy: 2.9 % / Rmerge F obs: 0.089 |
| Reflection shell | *PLUS Rmerge(I) obs: 0.344 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT / Resolution: 1.8→6 Å / Cross valid method: RFREE / σ(F): 1 / σ(I): 1 / Stereochemistry target values: Engh & Huber MOLECULAR REPLACEMENT / Resolution: 1.8→6 Å / Cross valid method: RFREE / σ(F): 1 / σ(I): 1 / Stereochemistry target values: Engh & Huber
| |||||||||||||||||||||||||
| Displacement parameters | Biso mean: -0.547 Å2
| |||||||||||||||||||||||||
| Refine analyze |
| |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→6 Å
| |||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.8→1.88 Å / Rfactor Rfree error: 0.01
| |||||||||||||||||||||||||
| Refinement | *PLUS % reflection Rfree: 5 % / Rfactor Rfree : 0.259 / Rfactor Rwork : 0.259 / Rfactor Rwork : 0.236 : 0.236 | |||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller












 PDBj
PDBj




















