[English] 日本語
 Yorodumi
Yorodumi- PDB-1l2j: Human Estrogen Receptor beta Ligand-binding Domain in Complex wit... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1l2j | ||||||
|---|---|---|---|---|---|---|---|
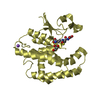
| Title | Human Estrogen Receptor beta Ligand-binding Domain in Complex with (R,R)-5,11-cis-diethyl-5,6,11,12-tetrahydrochrysene-2,8-diol | ||||||
 Components Components | ESTROGEN RECEPTOR BETA | ||||||
 Keywords Keywords | transcription receptor /  nuclear receptor / nuclear receptor /  transcription factor / transcription factor /  estrogen / estrogen /  antagonist antagonist | ||||||
| Function / homology |  Function and homology information Function and homology information receptor antagonist activity / nuclear steroid receptor activity / nuclear estrogen receptor activity / intracellular estrogen receptor signaling pathway / cellular response to estrogen stimulus / receptor antagonist activity / nuclear steroid receptor activity / nuclear estrogen receptor activity / intracellular estrogen receptor signaling pathway / cellular response to estrogen stimulus /  estrogen response element binding / estrogen response element binding /  steroid binding / ESR-mediated signaling / cellular response to estradiol stimulus / negative regulation of cell growth ... steroid binding / ESR-mediated signaling / cellular response to estradiol stimulus / negative regulation of cell growth ... receptor antagonist activity / nuclear steroid receptor activity / nuclear estrogen receptor activity / intracellular estrogen receptor signaling pathway / cellular response to estrogen stimulus / receptor antagonist activity / nuclear steroid receptor activity / nuclear estrogen receptor activity / intracellular estrogen receptor signaling pathway / cellular response to estrogen stimulus /  estrogen response element binding / estrogen response element binding /  steroid binding / ESR-mediated signaling / cellular response to estradiol stimulus / negative regulation of cell growth / positive regulation of DNA-binding transcription factor activity / Nuclear Receptor transcription pathway / Constitutive Signaling by Aberrant PI3K in Cancer / steroid binding / ESR-mediated signaling / cellular response to estradiol stimulus / negative regulation of cell growth / positive regulation of DNA-binding transcription factor activity / Nuclear Receptor transcription pathway / Constitutive Signaling by Aberrant PI3K in Cancer /  nuclear receptor activity / PIP3 activates AKT signaling / cell-cell signaling / PI5P, PP2A and IER3 Regulate PI3K/AKT Signaling / Extra-nuclear estrogen signaling / DNA-binding transcription factor activity, RNA polymerase II-specific / RNA polymerase II cis-regulatory region sequence-specific DNA binding / intracellular membrane-bounded organelle / nuclear receptor activity / PIP3 activates AKT signaling / cell-cell signaling / PI5P, PP2A and IER3 Regulate PI3K/AKT Signaling / Extra-nuclear estrogen signaling / DNA-binding transcription factor activity, RNA polymerase II-specific / RNA polymerase II cis-regulatory region sequence-specific DNA binding / intracellular membrane-bounded organelle /  chromatin / regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II / positive regulation of DNA-templated transcription / chromatin / regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II / positive regulation of DNA-templated transcription /  enzyme binding / negative regulation of transcription by RNA polymerase II / enzyme binding / negative regulation of transcription by RNA polymerase II /  signal transduction / positive regulation of transcription by RNA polymerase II / signal transduction / positive regulation of transcription by RNA polymerase II /  mitochondrion / mitochondrion /  DNA binding / zinc ion binding / DNA binding / zinc ion binding /  nucleoplasm / nucleoplasm /  nucleus nucleusSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.95 Å MOLECULAR REPLACEMENT / Resolution: 2.95 Å | ||||||
 Authors Authors | Shiau, A.K. / Barstad, D. / Radek, J.T. / Meyers, M.J. / Nettles, K.W. / Katzenellenbogen, B.S. / Katzenellenbogen, J.A. / Agard, D.A. / Greene, G.L. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 2002 Journal: Nat.Struct.Biol. / Year: 2002Title: Structural characterization of a subtype-selective ligand reveals a novel mode of estrogen receptor antagonism. Authors: Shiau, A.K. / Barstad, D. / Radek, J.T. / Meyers, M.J. / Nettles, K.W. / Katzenellenbogen, B.S. / Katzenellenbogen, J.A. / Agard, D.A. / Greene, G.L. #1:  Journal: Embo J. / Year: 1999 Journal: Embo J. / Year: 1999Title: Structure of the Ligand-binding Domain of Oestrogen Receptor Beta in the Presence of a Partial Agonist and a Full Antagonist Authors: Pike, A.C.W. / Brzozowski, A.M. / Hubbard, R.E. / Bonn, T. / Thorsell, A.G. / Engstrom, O. / Ljunggren, J. / Gustafsson, J.A. / Carlquist, M. #2:  Journal: J.Med.Chem. / Year: 1999 Journal: J.Med.Chem. / Year: 1999Title: Estrogen Receptor Subtype-selective Ligands: Asymmetric Synthesis and Biological Evaluation of cis- and trans-5,11-dialkyl-5,6,11,12-tetrahydrochrysenes Authors: Meyers, M.J. / Sun, J. / Carlson, K.E. / Katzenellenbogen, B.S. / Katzenellenbogen, J.A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1l2j.cif.gz 1l2j.cif.gz | 101.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1l2j.ent.gz pdb1l2j.ent.gz | 76.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1l2j.json.gz 1l2j.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/l2/1l2j https://data.pdbj.org/pub/pdb/validation_reports/l2/1l2j ftp://data.pdbj.org/pub/pdb/validation_reports/l2/1l2j ftp://data.pdbj.org/pub/pdb/validation_reports/l2/1l2j | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1l2iC  1a52S  1ereS  1errS  3erdS  3ertS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| Unit cell |
| ||||||||
| Details | The biological assembly is a homodimer which is not observed in the crystal. |
- Components
Components
| #1: Protein |  / ER-BETA / OESTROGEN RECEPTOR BETA / ER-BETA / OESTROGEN RECEPTOR BETAMass: 30571.119 Da / Num. of mol.: 2 / Fragment: ligand-binding domain (residues 256-505) Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Plasmid: pET15b / Production host: Homo sapiens (human) / Plasmid: pET15b / Production host:   Escherichia coli (E. coli) / Strain (production host): BL21(DE3)pLysS / References: UniProt: Q92731 Escherichia coli (E. coli) / Strain (production host): BL21(DE3)pLysS / References: UniProt: Q92731#2: Chemical |  (R,R)-Tetrahydrochrysene (R,R)-Tetrahydrochrysene#3: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.99 Å3/Da / Density % sol: 67.41 % | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Temperature: 295 K / Method: vapor diffusion, hanging drop / pH: 4.5 Details: 1.5-1.75 M Ammonium sulfate, 100 mM Sodium Acetate pH 4.8-5.2 294-296 K, pH 4.5, VAPOR DIFFUSION, HANGING DROP, temperature 295K | ||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 21-23 ℃ / PH range low: 5.2 / PH range high: 4.8 | ||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 103 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ALS ALS  / Beamline: 5.0.2 / Wavelength: 1.07 Å / Beamline: 5.0.2 / Wavelength: 1.07 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Feb 14, 1999 |
| Radiation | Monochromator: Double crystal / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.07 Å / Relative weight: 1 : 1.07 Å / Relative weight: 1 |
| Reflection | Resolution: 2.95→49.6 Å / Num. all: 14895 / Num. obs: 14895 / % possible obs: 99.7 % / Observed criterion σ(I): -3 / Redundancy: 4.3 % / Biso Wilson estimate: 46.7 Å2 / Rsym value: 0.045 / Net I/σ(I): 23.3 |
| Reflection shell | Resolution: 2.95→3.06 Å / Mean I/σ(I) obs: 3.4 / Num. unique all: 1507 / Rsym value: 0.302 / % possible all: 100 |
| Reflection | *PLUS Num. measured all: 64540 / Rmerge(I) obs: 0.045 |
| Reflection shell | *PLUS % possible obs: 100 % / Rmerge(I) obs: 0.302 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRIES 1A52, 1ERE, 1ERR, 3ERD, 3ERT Resolution: 2.95→49.6 Å / Rfactor Rfree error: 0.011 / Data cutoff high absF: 10000 / Isotropic thermal model: Restrained / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber Details: The structure was refined against data which had been sharpened with a -55 A**2 correction factor.
| ||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: flat model / Bsol: 33.127 Å2 / ksol: 0.333695 e/Å3 | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 40.08 Å2
| ||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.95→49.6 Å
| ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS | NCS model details: constrained | ||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.95→3.06 Å / Rfactor Rfree error: 0.044 / Total num. of bins used: 10
| ||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.259 / Rfactor Rfree : 0.299 / Rfactor Rwork : 0.299 / Rfactor Rwork : 0.259 : 0.259 | ||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| ||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor Rfree: 0.351 / Rfactor Rwork: 0.34 / Rfactor obs: 0.34 |
 Movie
Movie Controller
Controller












 PDBj
PDBj



