+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ije | ||||||
|---|---|---|---|---|---|---|---|
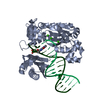
| Title | Nucleotide Exchange Intermediates in the eEF1A-eEF1Ba Complex | ||||||
 Components Components |
| ||||||
 Keywords Keywords |  TRANSLATION / TRANSLATION /  protein complex protein complex | ||||||
| Function / homology |  Function and homology information Function and homology informationEukaryotic Translation Elongation / eukaryotic translation elongation factor 1 complex / negative regulation of actin filament bundle assembly / HSF1 activation /  melatonin binding / regulation of translational termination / tRNA export from nucleus / melatonin binding / regulation of translational termination / tRNA export from nucleus /  Protein methylation / fungal-type vacuole membrane / translational elongation ...Eukaryotic Translation Elongation / eukaryotic translation elongation factor 1 complex / negative regulation of actin filament bundle assembly / HSF1 activation / Protein methylation / fungal-type vacuole membrane / translational elongation ...Eukaryotic Translation Elongation / eukaryotic translation elongation factor 1 complex / negative regulation of actin filament bundle assembly / HSF1 activation /  melatonin binding / regulation of translational termination / tRNA export from nucleus / melatonin binding / regulation of translational termination / tRNA export from nucleus /  Protein methylation / fungal-type vacuole membrane / translational elongation / actin filament bundle assembly / Protein methylation / fungal-type vacuole membrane / translational elongation / actin filament bundle assembly /  translation elongation factor activity / Neutrophil degranulation / cellular response to amino acid starvation / guanyl-nucleotide exchange factor activity / negative regulation of protein phosphorylation / maintenance of translational fidelity / negative regulation of protein kinase activity / GDP binding / translation elongation factor activity / Neutrophil degranulation / cellular response to amino acid starvation / guanyl-nucleotide exchange factor activity / negative regulation of protein phosphorylation / maintenance of translational fidelity / negative regulation of protein kinase activity / GDP binding /  actin filament binding / actin filament binding /  ribosome binding / ribosome binding /  cytoskeleton / cytoskeleton /  ribosome / ribosome /  translation / translation /  GTPase activity / GTP binding / GTPase activity / GTP binding /  protein kinase binding / protein kinase binding /  mitochondrion / mitochondrion /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Saccharomyces cerevisiae (brewer's yeast) Saccharomyces cerevisiae (brewer's yeast) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.4 Å MOLECULAR REPLACEMENT / Resolution: 2.4 Å | ||||||
 Authors Authors | Andersen, G.R. / Valente, L. / Pedersen, L. / Kinzy, T.G. / Nyborg, J. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 2001 Journal: Nat.Struct.Biol. / Year: 2001Title: Crystal structures of nucleotide exchange intermediates in the eEF1A-eEF1Balpha complex. Authors: Andersen, G.R. / Valente, L. / Pedersen, L. / Kinzy, T.G. / Nyborg, J. #1:  Journal: Mol.Cell / Year: 2000 Journal: Mol.Cell / Year: 2000Title: Structural basis for nucleotide exchange and competition with tRNA in the yeast elongation factor complex eEF1A:eEF1Balpha Authors: Andersen, G.R. / Pedersen, L. / Valente, L. / Chatterjee, I.I. / Kinzy, T.G. / Kjeldgaard, M. / Nyborg, J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ije.cif.gz 1ije.cif.gz | 118.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ije.ent.gz pdb1ije.ent.gz | 88.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ije.json.gz 1ije.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ij/1ije https://data.pdbj.org/pub/pdb/validation_reports/ij/1ije ftp://data.pdbj.org/pub/pdb/validation_reports/ij/1ije ftp://data.pdbj.org/pub/pdb/validation_reports/ij/1ije | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1g7cC  1ijfC  1f60S S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 50110.621 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Saccharomyces cerevisiae (brewer's yeast) / References: UniProt: P02994 Saccharomyces cerevisiae (brewer's yeast) / References: UniProt: P02994 |
|---|---|
| #2: Protein | Mass: 10046.277 Da / Num. of mol.: 1 / Fragment: catalytical C-terminal fragment Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Saccharomyces cerevisiae (brewer's yeast) Saccharomyces cerevisiae (brewer's yeast)Gene: TEF5 / Plasmid: pET11d / Production host:   Escherichia coli (E. coli) / Strain (production host): 834 DE3 / References: UniProt: P32471 Escherichia coli (E. coli) / Strain (production host): 834 DE3 / References: UniProt: P32471 |
| #3: Chemical | ChemComp-GDP /  Guanosine diphosphate Guanosine diphosphate |
| #4: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.21 Å3/Da / Density % sol: 44.35 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Temperature: 277 K / Method: vapor diffusion, sitting drop / pH: 8.2 Details: mmE2K, Tris, Hepes, KCl, DTT, pH 8.2, VAPOR DIFFUSION, SITTING DROP, temperature 277K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS pH: 7.2 Details: Pedersen, L.P., (2000) Acta Crystallogr., D57, 159. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: ENRAF-NONIUS FR571 / Wavelength: 1.5418 Å ROTATING ANODE / Type: ENRAF-NONIUS FR571 / Wavelength: 1.5418 Å |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Jan 13, 2001 / Details: graphite monochromator |
| Radiation | Monochromator: graphite / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.5418 Å / Relative weight: 1 : 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.4→20 Å / Num. all: 21448 / Num. obs: 21223 / % possible obs: 100 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 4.2 % / Biso Wilson estimate: 22 Å2 / Rmerge(I) obs: 0.064 / Net I/σ(I): 22.1 |
| Reflection shell | Resolution: 2.4→2.49 Å / Redundancy: 4.2 % / Rmerge(I) obs: 0.205 / Num. unique all: 2110 |
| Reflection | *PLUS Lowest resolution: 20 Å / % possible obs: 100 % |
| Reflection shell | *PLUS Highest resolution: 2.4 Å / % possible obs: 99 % / Mean I/σ(I) obs: 6.8 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: pdb entry 1f60 Resolution: 2.4→19.88 Å / Rfactor Rfree error: 0.007 / Data cutoff high absF: 346790.09 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / Stereochemistry target values: Engh and Huber
| ||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 20.68 Å2 / ksol: 0.335 e/Å3 | ||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 23.1 Å2
| ||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.4→19.88 Å
| ||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.4→2.55 Å / Rfactor Rfree error: 0.021 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Version: 1 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS σ(F): 0 / % reflection Rfree: 5.4 % / Rfactor Rfree : 0.25 : 0.25 | ||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 23.1 Å2 | ||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| ||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor Rfree: 0.276 / % reflection Rfree: 5.1 % / Rfactor Rwork: 0.228 |
 Movie
Movie Controller
Controller












 PDBj
PDBj


















